Abstract
Atypical fibroxanthomas and pleomorphic dermal sarcomas are tumors arising in sun-damaged skin of elderly patients. They have differing prognoses and are currently distinguished using histological criteria, such as invasion of deeper tissue structures, necrosis and lymphovascular or perineural invasion. To investigate the as-yet poorly understood genetics of these tumors, 41 atypical fibroxanthomas and 40 pleomorphic dermal sarcomas were subjected to targeted next-generation sequencing approaches as well as DNA copy number analysis by comparative genomic hybridization. In an analysis of the entire coding region of 341 oncogenes and tumor suppressor genes in 13 atypical fibroxanthomas using an established hybridization-based next-generation sequencing approach, we found that these tumors harbor a large number of mutations. Gene alterations were identified in more than half of the analyzed samples in FAT1, NOTCH1/2, CDKN2A, TP53, and the TERT promoter. The presence of these alterations was verified in 26 atypical fibroxanthoma and 35 pleomorphic dermal sarcoma samples by targeted amplicon-based next-generation sequencing. Similar mutation profiles in FAT1, NOTCH1/2, CDKN2A, TP53, and the TERT promoter were identified in both atypical fibroxanthoma and pleomorphic dermal sarcoma. Activating RAS mutations (G12 and G13) identified in 3 pleomorphic dermal sarcoma were not found in atypical fibroxanthoma. Comprehensive DNA copy number analysis demonstrated a wide array of different copy number gains and losses, with similar profiles in atypical fibroxanthoma and pleomorphic dermal sarcoma. In summary, atypical fibroxanthoma and pleomorphic dermal sarcoma are highly mutated tumors with recurrent mutations in FAT1, NOTCH1/2, CDKN2A, TP53, and the TERT promoter, and a range of DNA copy number alterations. These findings suggest that atypical fibroxanthomas and pleomorphic dermal sarcomas are genetically related, potentially representing two ends of a common tumor spectrum and distinguishing these entities is at present still best performed using histological criteria.
Similar content being viewed by others
Main
Atypical fibroxanthoma and pleomorphic dermal sarcoma are rare mesenchymal tumors typically arising in sun-damaged skin of elderly patients. The pathogenesis of these tumors is not well understood.
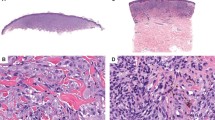
Atypical fibroxanthoma are tumors that occur in sun-damaged skin, primarily in the head and neck region of elderly patients.1, 2, 3, 4 More frequent in males, these tumors are often well circumscribed, rapidly growing, and demonstrate an exophytic growth pattern. Known risk factors include UV exposure, irradiation, xeroderma pigmentosum, and organ transplantation.5 Histologically, the tumors are composed of atypical spindled and pleomorphic tumor cells, including tumor giant cells. Primarily located within the dermis, tumors may have limited extension into the subcutis. High mitotic activity is customary, with presentation of frequent atypical mitotic figures.2 Atypical fibroxanthoma do not invade the deep soft tissue, and despite increased proliferative activity, histological features such as necrosis, and lymphovascular and/or perineural invasion are not observed. A diagnosis of atypical fibroxanthoma requires exclusion of other neoplasms, in particular, melanoma, squamous cell carcinoma, and leiomyosacroma. Atypical fibroxanthoma generally has a good prognosis, and usually complete excision and regular follow-up are recommended.1
Pleomorphic dermal sarcomas demonstrate similar morphology to atypical fibroxanthoma, but present more aggressive histological features such as extensive involvement of the subcutis and/or deeper structures, areas of tumor necrosis, and lymphovascular or perineural invasion.6 Pleomorphic dermal sarcoma display more aggressive clinical behavior than atypical fibroxanthoma and are categorized as tumors with low-grade malignant potential based on their potential for local recurrence and metastasis.7 The tumors now defined as pleomorphic dermal sarcoma have also been referred to as cutaneous undifferentiated pleomorphic sarcomas or as superficial malignant fibrous histiocytomas in the past.8, 9, 10 There remains a lack of effective therapies for metastasized pleomorphic dermal sarcoma.
Distinguishing atypical fibroxanthoma from pleomorphic dermal sarcoma is critical due to their differences in clinical behavior. Discriminating these entities based on cell morphology alone is not possible as their cytologic features are similar. Histological criteria applied to distinguish pleomorphic dermal sarcoma from atypical fibroxanthoma are the following: larger tumor size; extensive infiltration of subcutis, invasion of fascia or muscle; necrosis; and vascular or perineural invasion.2, 6 Attempts to differentiate between atypical fibroxanthoma and pleomorphic dermal sarcoma based on biopsy specimens should be avoided as these may not demonstrate the deepest extent of tumor involvement or miss histological criteria such as necrosis or vascular/perineural invasion present in other regions of the tumor. Attempts to identify immunohistochemical markers facilitating the distinction of atypical fibroxanthoma from pleomorphic dermal sarcoma (eg, CD99 and LN-2 (refs 11, 12) have not proven useful in routine practice.1, 2, 3, 13
Little is known of the genetic events leading to the development of atypical fibroxanthoma and pleomorphic dermal sarcoma. Previous smaller studies identified UV-signature mutations in TP53 in atypical fibroxanthoma (7/10 (ref. 14) and 4/6 (ref. 15) cases) and pleomorphic dermal sarcoma (1/4 cases, diagnosed as ‘malignant fibrous histiocytoma’15), as well as one HRAS and one KRAS mutation in 8 pleomorphic dermal sarcoma (diagnosed as ‘malignant fibrous histiocytoma’) analyzed.16 In another study, pleomorphic dermal sarcoma (diagnosed as ‘undifferentiated pleomorphic sarcoma‘) were found to harbor more frequent DNA copy number alterations than atypical fibroxanthoma.17 We have reported high frequencies of TERT promoter mutations in atypical fibroxanthoma and pleomorphic dermal sarcoma.18 A recent study of 5 atypical fibroxanthoma and 5 pleomorphic dermal sarcoma demonstrated frequent TP53 mutations, as well as individual CDKN2A, HRAS, KNSTRN, and PIK3CA gene mutations.19
The goal of our study was to try and identify recurrent gene mutations in atypical fibroxanthoma and pleomorphic dermal sarcoma applying newer sequencing technologies and genome-wide DNA copy number analysis. In addition to generally gaining a better understanding of the pathogenesis of these tumor entities, we attempted to determine whether genetic alterations may be an additional diagnostic aid in distinguishing these tumors.
Materials and methods
Sample Selection
Atypical fibroxanthoma and pleomorphic dermal sarcoma tumor samples were obtained from the tissue archives of Dermatopathology Duisburg (24 atypical fibroxanthoma) and Dermatopathology Friedrichshafen (17 atypical fibroxanthoma and 20 pleomorphic dermal sarcoma), Germany, as well as from the Department of Pathology, Western General Hospital, The University of Edinburgh, Edinburgh, UK (20 pleomorphic dermal sarcoma, which were previously described by Miller et al.7). The study was done in accordance with the guidelines set forth by the ethics committee of the University of Duisburg-Essen.
Histopathology and Immunohistochemistry
Histologic sections of all tumors were reviewed, and the diagnoses confirmed by at least two dermatopathologists (TB, TM, KGG, and JS). Available clinical and pathologic data were analyzed, including age, sex, site, size, depth, polyploid architecture, tumor mitotic rate, ulceration, necrosis, subcutis invasion, smooth muscle or fascia invasion, infiltrative or pushing border, lympovascular invasion, and perineural invasion. Immunohistochemical markers applied to all tumors, including the following: pan-cytokeratin (MNF116; 1/500, AE1/AE3; 1/50); CD31 (JC70A; 1/100) or CD34 (QBEnd-10; 1/50); S100 (polyclonal; 1/2000); desmin (D33; 1/100); smooth muscle actin (ASMA 1A4; 1/500); and Melan-A (A103; 1/1000). The described antibodies were obtained from DAKO Hamburg/Germany (Duisburg and Friedrichshafen cases). Stains in the cases from Edinburgh were performed as described previously.7
DNA Isolation
The 10 μm-thick sections were cut from formalin-fixed, paraffin-embedded tumor tissues. Sections were deparaffinized and tumor-bearing areas manually macrodissected according to routine procedures. Isolation of genomic DNA was performed applying the QIAamp DNA Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions.
DNA Copy Number Analysis
Array-based comparative genomic hybridization was used to perform analysis of DNA copy number aberrations. The methods for hybridization and analysis, including GISTIC 2.0 statistical analysis, have been described previously.20, 21, 22, 23 In individual cases whole-genome amplification was performed using Sigma’s GenomePlex Single Cell Whole Genome Amplification Kit as described previously.24
Hybridization-Capture-Based Next-Generation Sequencing for Known Oncogene Mutations
Custom DNA probes were designed for targeted sequencing of all exons and selected introns of 341 oncogenes and tumor suppressor genes (MSK-IMPACT assay).25 Briefly, genomic DNA from tumor samples was used to prepare barcoded libraries using the KAPA HTP protocol (Kapa Biosystems, Wilmington, MA, USA) and the Biomek FX system (Beckman Coulter, Brea, CA, USA). Libraries were pooled, captured, and subsequently sequenced on an Illumina HiSeq 2500 system as paired-end reads. Sequenced reads were trimmed to remove vestigial adaptor sequences using TrimGalore,26 and were aligned to the hg19 human reference genome using BWA.27 PCR duplicates were removed from the alignment output, and the aligned reads were subjected to local indel realignment and base quality recalibration using GATK.28 Somatic variant calling was performed using MuTect29 for single-nucleotide variants (SNVs) and SomaticIndelDetector28 for indels. Significant copy number gains and losses were detected by requiring a greater-than twofold change in normalized coverage between tumor and a comparator reference fresh-frozen paraffin-embedded normal. Somatic structural variants were detected using DELLY,30 requiring both paired-end and split-read support.
Amplicon-Based Targeted Next-Generation Sequencing (Amplicon Next-Generation Sequencing)
A custom amplicon-based sequencing panel covering 11 genes (HRAS, KRAS, NRAS, CDKN2A, FAT1, KNSTRN, TP53, NOTCH1, NOTCH2, PIK3CA, and TSC2) was designed (Table 1). This panel included the genes most frequently found mutated in atypical fibroxanthoma in our prior screen (impact next-generation sequencing), as well as the RAS genes, which were previously reported to be mutated in pleomorphic dermal sarcomas.16 Library preparation was performed applying the GeneRead Library Prep Kit from QIAGEN according to the manufacturer’s instructions. NEBNext Ultra DNA Library Prep Mastermix Set and NEBNext Multiplex Oligos for Illumina from New England Biolabs were used for adapter ligation and barcoding individual samples. Sequencing was performed on an Illumina MiSeq next-generation sequencer, sequencing up to 24 samples in parallel. Sequencing the sample cohort, an average coverage of 2803 reads was achieved with 82% of the target area having a minimum coverage of 30 reads.
Sequence Analysis
CLC Cancer Research Workbench from QIAGEN was applied as previously reported31 for sequence analysis. Briefly, the CLC workflow included adapter trimming and read pair merging before mapping to the human reference genome (hg19). Subsequently, insertions and deletions as well as SNV detection, local realignment, and primer trimming followed. Various databases were cross-referenced (COSMIC, ClinVar, dbSNP, 1000 Genomes Project, HAPMAP, and PhastCons-Conservation_scores_hg19) regarding information on mutation type, single-nucleotide polymorphisms, and conservation scores. Csv files were further analyzed manually screening for protein-coding mutations predicted to result in non-synonymous amino-acid changes. Mutations were considered if the overall coverage of the mutation site was ≥30 reads, ≥10 reads reported the mutated variant and the frequency of mutated reads was ≥10%.
Direct (Sanger) Sequencing
PCR amplification of the TERT promoter region was done as previously described.18 PCR products were purified with the QIAquick PCR Purification Kit (Qiagen) and used as templates for sequencing. The sequencing chromatogram files were examined using Chromas software (version 2.01, University of Sussex, Brighton, UK) or Sequencher software (version 5.1, Gene Codes Corporation, Ann Arbor, MI, USA).
Associations of Gene Mutation Status with Clinical and Pathologic Parameters
We investigated associations of mutation status with available clinical and pathological parameters using non-parametric tests, χ2-tests, and Fisher exact tests as appropriate. All statistical analyses were performed using PASW Statistics 18 software (International Business Machines, Armonk, NY, USA). A P-value of ≤0.05 was considered statistically significant.
Results
Study Cohort
In all, 81 (41 atypical fibroxanthoma and 40 pleomorphic dermal sarcoma) tumors were analyzed, of which 77 were primary tumor samples and 4 were recurrent pleomorphic dermal sarcoma. A total of 67 patients were male and 14 were female. The median age at diagnosis was 82 (range 47–102) years overall; 79 (range 65–93) years for atypical fibroxanthoma and 84 (range 47–102) years for pleomorphic dermal sarcoma. An overview of the different genetic analysis applied to each sample is given in Supplementary Table 1.
Clinical and Pathologic Features of Atypical Fibroxanthoma and Pleomorphic Dermal Sarcoma
Clinical and pathologic features are listed in Table 1. Tumors were negative for most listed immunohistochemical markers. Focal or patchy smooth muscle actin expression was seen in 14 (34%) atypical fibroxanthoma and 11 (28%) pleomorphic dermal sarcoma. As previously reported,7 10 (25%) pleomorphic dermal sarcoma cases showed focal expression of CD31 and 1 pleomorphic dermal sarcoma exhibited limited aberrant expression of Melan-A (S100 was negative).
Genomic Findings
Hybridization-capture-based next-generation sequencing for known oncogene mutations
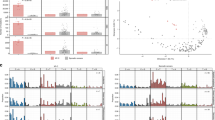
A considerable number of mutations were identified in the 13 atypical fibroxanthoma tumor samples analyzed with an overall mean sequencing depth of 492-fold. The average number of mutations identified in each sample was 55, the range was between 34 and 94 mutations. Genes mutated in more than 3 tumors are demonstrated in Figure 1 (all mutations identified are listed in Supplementary Table 2). Particularly frequent were TP53 mutations, identified in all samples, with 10/13 (77%) samples harboring inactivating mutations. TERT promoter mutations were identified in 12/13 (92%) samples. In FAT1, 8 of 13 (62%) samples harbored mutations, 7 (87.5%) were inactivating. Other apparent alterations affected CDKN2A with 6 of 13 (46%) mutations, 5 (83%) inactivating. NOTCH1 was mutated in 11 of 13 (85%) samples, also with 5 (45%) inactivating mutations (5). Although the panel covers most known activating mutations in human cancer,25 no recurrent known functionally activating protein-coding mutations were identified in these tumors.
Distribution of mutations in atypical fibroxanthomas. Demonstrated are data from 13 atypical fibroxanthomas sequenced with a hybridization-based screen covering 341 oncogenes and tumor suppressors. All exons of the genes were analyzed. The genes presented are those found to have been mutated in at least 3 samples. The mutation type is according to the legend. No mutation identified (wild type) is signified by no fill, missense mutations of unclear consequence by gray filled boxes, loss-of-function (non-sense or frameshift) mutations are signified by red filled boxes, and activating mutations in the TERT promoter by blue filled boxes.
Targeted amplicon-based next-generation sequencing (amplicon next-generation sequencing)
To further assess and validate the distribution of many of the identified recurrent mutations a custom panel was designed and applied covering the most frequently mutated genes (Table 1). In addition to genes identified in the described screen for oncogene mutations, the RAS genes, previously reported to be mutated in pleomorphic dermal sarcoma,16 and KNSTRN, a gene reported mutated in squamous cell carcinoma,32 were included in the panel. Overall, 26 atypical fibroxanthoma and 35 pleomorphic dermal sarcoma were sequenced. The mutation profiles of both tumor entities were similar (Figure 2; Supplementary Table 3). High numbers of TP53 mutations in 22/26 (85%) atypical fibroxanthoma and 31/35 (89%) pleomorphic dermal sarcoma, NOTCH1 mutations in 14/26 (54%) atypical fibroxanthoma and 22/35 (63%) pleomorphic dermal sarcoma, NOTCH2 mutations in 11/26 (42%) atypical fibroxanthoma, and 17/35 (49%) pleomorphic dermal sarcoma, as well as FAT1 mutations in 15/26 (58%) atypical fibroxanthoma and 24/35 (69%) pleomorphic dermal sarcoma were identified. Less frequently detected were TSC2, PIK3CA, and CDKN2A mutations (Figure 2).
UV-signature mutation analysis
To assess the role of UV exposure in the pathogenesis of atypical fibroxanthoma and pleomorphic dermal sarcoma, an analysis of mutations identified in the TP53 gene was performed, assessing the amount of potentially UV-induced mutations. An approach that has been previously utilized for cutaneous tumors.14, 33, 34, 35 In 61 tumors, 93 mutations in TP53 were identified, 37 in 25 atypical fibroxanthoma and 56 in 36 pleomorphic dermal sarcoma. The frequency of mutations with a UV signature was high in both tumors, with C>T (G>A) and CC>TT (GG>AA) mutations being detected in 43% (16/37) and 14% (5/37) of mutations in atypical fibroxanthoma and 57% (32/56) and 7% (4/56) of mutations in pleomorphic dermal sarcoma, respectively.
TERT promoter
The TERT promoter was analyzed by Sanger sequencing as previously described.18 The majority of tumors 19/20 (95%) atypical fibroxanthoma and 18/24 (75%) pleomorphic dermal sarcoma where sequencing was successful, demonstrated at least one known activating hot-spot mutation in the TERT promoter.
Copy number alterations
Copy number information was performed by comparative genomic hybridization in 42 samples, 20 atypical fibroxanthoma, and 22 pleomorphic dermal sarcoma samples. The overall results are demonstrated in Figure 3. Tumors showed a considerable number of alterations. Evident differences in terms of alteration profiles between atypical fibroxanthoma and pleomorphic dermal sarcoma were not apparent. Common were losses of 8p, 9p, and 9q. Larger deletions involving most of Chr. 13, 16, and 18 were also apparent. Gains were less frequent however identified in Chr. 1q, 8q, 17q, and 19p. The profile of alterations was very similar in atypical fibroxanthoma and pleomorphic dermal sarcoma (Figure 3).
Copy number profiles in atypical fibroxanthomas and pleomorphic dermal sarcomas. Shown here are the comparative genomic hybridization results of 20 atypical fibroxanthoma shown on top, 22 pleomorphic dermal sarcomas shown in the middle panel, and all 42 samples together, shown on the bottom panel. All groups were analyzed identically with Agilent software. Alterations are displayed as penetrance plots, gains in red, losses in green.
Associations of clinicopathologic and mutation status in atypical fibroxanthoma and pleomorphic dermal sarcoma
The complete analysis is presented in Table 2. Histological criteria used to distinguish atypical fibroxanthoma from pleomorphic dermal sarcoma were found to be clearly associated with tumor type, including the following: necrosis (P<0.001); invasion into subcutis (P<0.001); invasion into fascia/skeletal muscle (<0.001); border (P<0.001); lymphovascular invasion (P=0.001); and perineural invasion (P<0.001). Differences in terms of gene mutation status between atypical fibroxanthoma and pleomorphic dermal sarcoma demonstrated in Table 3 were not found to be statistically significant. An analysis of gene mutation status with individual histopathological criteria (Supplementary Table 4) demonstrated no correlations with mutation status with the exception of age with NOTCH2 (P=0.05), tumor depth with TP53 (P=0.05), and necrosis with TERT promoter (P=0.002) mutation status (shown in detail in Supplementary Table 4).
Discussion
The genetic underpinnings of atypical fibroxanthoma and pleomorphic dermal sarcoma, as well as the relationship between these two tumor entities has been a question of debate for many years. We believe the findings of our study contribute to a better understanding of these entities, demonstrating that these tumors harbor a large number of common genetic alterations. In addition to known TERT promoter and TP53 mutations, alterations in CDKN2A, FAT1, NOTCH1, and NOTCH2 were identified particularly frequently.
The comprehensive screen for known cancer mutations in 341 known cancer genes demonstrated a very high frequency of mutations overall. A selection of these genes is shown in Figure 1 (only genes mutated in at least three tumors are included). Recurrent gene mutations known to activate signaling pathways such as the MAPK pathway were not identified. Neither were recurrent hot-spot mutations in genes other that the TERT promoter.18 What was striking is that in the large array of frequently mutated genes, some genes were mutated in a high percentage of samples, including frequent loss-of-function mutations.
TERT promoter mutations, originally identified in cutaneous melanoma36, 37 were subsequently identified in a wide array of cancers38 and previously shown by our group to occur in the majority (>70%) of atypical fibroxanthoma and pleomorphic dermal sarcoma samples.18 This finding was validated in the present study (Figures 1 and 2).
TP53 has been known to be altered in atypical fibroxanthoma and pleomorphic dermal sarcoma for more than two decades.14, 15 Our study validates this finding, demonstrating TP53 mutations in the majority of all tumor samples (atypical fibroxanthoma MSK-IMPACT next-generation sequencing=100%, atypical fibroxanthoma amplicon next-generation sequencing=89%, and pleomorphic dermal sarcoma amplicon next-generation sequencing=86%) with a considerable number of samples harboring inactivation mutations (atypical fibroxanthoma MSK-IMPACT next-generation sequencing=77%, atypical fibroxanthoma amplicon next-generation sequencing=36%, and pleomorphic dermal sarcoma amplicon next-generation sequencing=31%). In addition, losses of the TP53 gene locus on 17p did occur. These findings validate TP53 as a highly altered and important gene in atypical fibroxanthoma and pleomorphic dermal sarcoma.
FAT1 mutations were previously identified in various cancers, including glioblastoma, colorectal and head and neck cancer. Both losses and mutations were reported.39 In our data, both frequent losses of Chr. 4q (Figure 3) as well as a very high frequency of gene mutations were detected. In all screens performed, the mutation frequency was ~60% of tumors (atypical fibroxanthoma MSK-IMPACT next-generation sequencing=62%, atypical fibroxanthoma amplicon next-generation sequencing=58%, and pleomorphic dermal sarcoma amplicon next-generation sequencing=67%), the percentage of inactivating FAT1 mutations >40% (atypical fibroxanthoma impact next-generation sequencing=64%, atypical fibroxanthoma amplicon next-generation sequencing=42%, and pleomorphic dermal sarcoma amplicon next-generation sequencing=44%). FAT1 inactivation has been associated with increased β-catenin and Wnt signaling,39 suggesting Wnt signaling may play an important role in atypical fibroxanthoma and pleomorphic dermal sarcoma pathogenesis.
NOTCH signaling, having a critical role in tissue development, has long been known to have a relevant role in cancer pathogenesis.40, 41, 42 Originally associated with activating genetic alterations in hematological malignancies,43 recurrent loss-of-function alterations have also been identified in a wide range of malignancies, including head and neck and cutaneous squamous cell carcinoma.40, 44, 45, 46 Very similar to cutaneous squamous cell carcinoma, we identified the highest frequency of mutations in NOTCH1 (atypical fibroxanthoma MSK-IMPACT next-generation sequencing=85%, atypical fibroxanthoma amplicon next-generation sequencing=54%, and pleomorphic dermal sarcoma amplicon next-generation sequencing=66%) followed by NOTCH2 (atypical fibroxanthoma MSK-IMPACT next-generation sequencing=62%, atypical fibroxanthoma amplicon next-generation sequencing=42%, and pleomorphic dermal sarcoma amplicon next-generation sequencing=49%). Both genes also demonstrated a considerable amount of inactivating mutations (Figures 1 and 2). Chromosomal losses of the gene loci (NOTCH1 9q and NOTCH2 1p) were rare and probably are not of considerable relevance.
CDKN2A, which codes for p14 and p16, is one of the most frequently lost tumor suppressors in human neoplasias47 and is clearly also of relevance in atypical fibroxanthoma and pleomorphic dermal sarcoma. Not only was it found mutated in around half of the samples in the atypical fibroxanthoma impact next-generation sequencing screen (6 of 13=46%) but copy number analysis (comparative genomic hybridization) also demonstrated the locus on Chr. 9p to be lost in around half of atypical fibroxanthoma and pleomorphic dermal sarcoma tumor samples (Figure 3). The low mutation rate detected in the amplicon next-generation sequencing screen is most likely due to poor coverage of the gene in our sequencing panel (Table 1), and losses of the gene locus being more frequent than focal gene mutations.
The analysis of C>T (G>A) and CC>TT (GG>AA) mutations associated with UV exposure in the TP53 gene showed these were frequent both in atypical fibroxanthoma and pleomorphic dermal sarcoma, representing more than half of the identified alterations in both tumor entities (57% in atypical fibroxanthoma and 64% in pleomorphic dermal sarcoma). Interestingly, C>T (G>A) alterations were detected more frequently in pleomorphic dermal sarcoma than atypical fibroxanthoma (57% and 43% of mutations, respectively), however CC>TT (GG>AA) were more frequent in atypical fibroxanthoma than pleomorphic dermal sarcoma (14% and 7% of mutations, respectively). Whereas C>T alterations are frequently UV-induced, there association with UV exposure is not as strong as is the case for CC>TT alterations, which are considered to be virtually pathognomonic for UV exposure.33, 48 The overall high frequency of these mutations (both C>T and CC>TT) in both atypical fibroxanthoma and pleomorphic dermal sarcoma clearly supports UV exposure being a relevant pathogenic event in both of these tumor entities.
No associations between mutation status and diagnosis of atypical fibroxanthoma and pleomorphic dermal sarcoma were observed. The rare associations identified in our study with individual clinicopathological parameters: age with NOTCH2 mutation status (P=0.05); tumor depth and TP53 (P=0.05) mutation status; and necrosis and TERT promoter mutation status (P=0.002; Supplementary Table 4) will need to be validated in larger independent cohorts.
Alterations in many genes including those recurrently detected in atypical fibroxanthoma and pleomorphic dermal sarcoma (TP53, CDKN2A, FAT1, NOTCH1/2, and the TERT promoter) have been reported by various groups in cutaneous squamous cell carcinoma.32, 49, 50 Given that these tumors (cutaneous squamous cell carcinomas, atypical fibroxanthoma, and pleomorphic dermal sarcoma) all arise in areas of high sun exposure, it is likely that the similarities in mutation profiles are in part due to a common UV-induced pathogenesis of these entities. The signaling pathways essential for tumor development in these tumors may also be similar. While this is an intriguing hypothesis, additional genetic and functional studies will be required to fully reveal similarities and differences in the pathogenesis of these tumors.
One of the goals of our study was to try and identify genetic differences distinguishing atypical fibroxanthoma and pleomorphic dermal sarcoma. Our study finds little evidence for genetic differences between these entities. The copy number profiles determined here in a relatively large cohort of tumors (atypical fibroxanthoma=20 and pleomorphic dermal sarcoma=22) are very similar, allowing no apparent separation of these entities. Genetically both tumor groups harbored similar frequencies of FAT1, TP53, CDKN2A, TERT promoter, and NOTCH alterations. What is interesting is that activating RAS mutations, previously identified as being present in pleomorphic dermal sarcoma (diagnosed then as MFH) but not atypical fibroxanthoma, were also found to be distributed in a similar manner in our cohort. However, these activating mutations (2 HRAS G12S, G13V and 1 KRAS G12D) were rare, present only in 3 of 35 (9%) of pleomorphic dermal sarcoma. As such it seems unlikely determining RAS mutation status will be of use in clinical distinction between atypical fibroxanthoma and pleomorphic dermal sarcoma.
Considering the high mutation frequencies and copy number alterations present in both atypical fibroxanthoma and pleomorphic dermal sarcoma, our study suggests it will remain difficult to fully understand the key genetic mechanisms involved in the development of these tumors. Whole-exome or whole-genome sequencing may offer more insights, however distinguishing relevant driver mutations from the accompanying very high number of passenger mutations could prove a considerable challenge. To which extent alterations can be identified allowing a clear distinction of atypical fibroxanthoma and pleomorphic dermal sarcoma will remain to be seen.
As most mutations identified in our screen are assumed to be loss-of-function mutations, it may prove difficult identifying effective treatment strategies targeting cell intrinsic signaling pathways in atypical fibroxanthoma or pleomorphic dermal sarcoma. On the other hand, given that high mutational load is associated with better responses to immunotherapy with anti-CTLA4 and anti-PD1 immune checkpoint blockade therapies in many different cancer entities,51, 52, 53, 54 our findings may suggest immunotherapy as a promising therapeutic approach for recurrent or metastatic pleomorphic dermal sarcoma.
Weaknesses of our study are that only a selection of genes were analyzed. Whole-exome or whole-genome approaches should prove valuable in the future, however considering the high mutation frequency we observed, large numbers of tumors will need to be assessed to identify relevant recurrent events. Obtaining the necessary number of tissue samples (preferably fresh-frozen with paired germline DNA) and performing a meaningful bioinformatics analysis may prove challenging. The lack of available paired normal tissue in our study is another considerable caveat. Although most germline variants will have been successfully excluded by cross-referencing SNP databases, we cannot exclude the possibility that occasional germline variants were interpreted as somatic mutations. As a result, the frequency of somatic mutations we report is likely to be somewhat higher than is actually the case. However, the comparable results obtained in our study through two separate NGS approaches, capture (MSK-IMPACT NGS) and amplicon (amplicon NGS) and analyzed by two different bioinformatics approaches, supports the validity and accuracy of the somatic mutation profiles we report. However, larger future studies with paired germline DNA will be valuable to validate our findings and further elucidate the genetic landscape of these tumor entities.
Despite certain shortcomings, our study is the most comprehensive to date identifying both a number of previously unrecognized recurrent gene mutations (ie, NOTCH1, NOTCH2, and FAT1) and a fairly comprehensive picture of copy number alterations in these tumors. Our findings do not fully resolve the long-standing debate as to the relationship between atypical fibroxanthoma and pleomorphic dermal sarcoma. However, we do believe the similar gene mutations and copy number profiles identified argue strongly the tumors are related, potentially representing entities along a common tumor spectrum. Whereas future studies may manage to identify robust genetic or other biomarkers allowing a clear diagnostic distinction of these two tumor entities, our current findings suggest the established histological criteria for distinguishing these two entities (eg, larger tumor size with extensive involvement of the subcutis, musculature or fascia, perineural or perivascular invasion, or tissue necrosis) will remain of paramount importance in determining the prognosis and correct clinical management of affected patients.
In summary, our data demonstrate that atypical fibroxanthoma and pleomorphic dermal sarcoma harbor a large number of genetic alterations, including frequent TP53, CDKN2A, TERT promoter, NOTCH1 and NOTCH2, and FAT1 gene alterations, and a wide range of copy number alterations. Rare activating RAS mutations were only present in pleomorphic dermal sarcoma (9% of tumors). Whereas we believe these findings offer valuable insights into the pathogenesis of these tumors, it remains to be shown if they will be of diagnostic or therapeutic value in a clinical setting.
References
Iorizzo LJ 3rd, Brown MD . Atypical fibroxanthoma: a review of the literature. Dermatol Surg 2011;37:146–157.
Gru AA, Santa Cruz DJ . Atypical fibroxanthoma: a selective review. Semin Diagn Pathol 2013;30:4–12.
McCalmont TH . AFX: what we now know. J Cutan Pathol 2011;38:853–856.
Mentzel T, Requena L, Brenn T . Atypical fibroxanthoma revisited. Surg Pathol Clin 2017;10:319–335.
McCoppin HH, Christiansen D, Stasko T et al. Clinical spectrum of atypical fibroxanthoma and undifferentiated pleomorphic sarcoma in solid organ transplant recipients: a collective experience. Dermatol Surg 2012;38:230–239.
McCalmont TH . Correction and clarification regarding AFX and pleomorphic dermal sarcoma. J Cutan Pathol 2012;39:8.
Miller K, Goodlad JR, Brenn T . Pleomorphic dermal sarcoma: adverse histologic features predict aggressive behavior and allow distinction from atypical fibroxanthoma. Am J Surg Pathol 2012;36:1317–1326.
Ozzello L, Stout AP, Murray MR . Cultural characteristics of malignant histiocytomas and fibrous xanthomas. Cancer 1963;16:331–344.
Fletcher CD . The evolving classification of soft tissue tumours: an update based on the new WHO classification. Histopathology 2006;48:3–12.
Henderson MT, Hollmig ST . Malignant fibrous histiocytoma: changing perceptions and management challenges. J Am Acad Dermatol 2012;67:1335–1341.
Hartel PH, Jackson J, Ducatman BS et al. CD99 immunoreactivity in atypical fibroxanthoma and pleomorphic malignant fibrous histiocytoma: a useful diagnostic marker. J Cutan Pathol 2006;33:24–28.
Lazova R, Moynes R, May D et al. LN-2 (CD74). A marker to distinguish atypical fibroxanthoma from malignant fibrous histiocytoma. Cancer 1997;79:2115–2124.
Hanlon A, Stasko T, Christiansen D et al. LN2, CD10, and Ezrin do not distinguish between atypical fibroxanthoma and undifferentiated pleomorphic sarcoma or predict clinical outcome. Dermatol Surg 2017;43:431–436.
Dei Tos AP, Maestro R, Doglioni C et al. Ultraviolet-induced p53 mutations in atypical fibroxanthoma. Am J Pathol 1994;145:11–17.
Sakamoto A, Oda Y, Itakura E et al. Immunoexpression of ultraviolet photoproducts and p53 mutation analysis in atypical fibroxanthoma and superficial malignant fibrous histiocytoma. Mod Pathol 2001;14:581–588.
Sakamoto A, Oda Y, Itakura E et al. H-, K-, and N-ras gene mutation in atypical fibroxanthoma and malignant fibrous histiocytoma. Hum Pathol 2001;32:1225–1231.
Mihic-Probst D, Zhao J, Saremaslani P et al. CGH analysis shows genetic similarities and differences in atypical fibroxanthoma and undifferentiated high grade pleomorphic sarcoma. Anticancer Res 2004;24:19–26.
Griewank KG, Schilling B, Murali R et al. TERT promoter mutations are frequent in atypical fibroxanthomas and pleomorphic dermal sarcomas. Mod Pathol 2013;27:502–508.
Helbig D, Ihle MA, Putz K et al. Oncogene and therapeutic target analyses in atypical fibroxanthomas and pleomorphic dermal sarcomas. Oncotarget 2016;7:21763–21774.
Wiesner T, Obenauf AC, Murali R et al. Germline mutations in BAP1 predispose to melanocytic tumors. Nat Genet 2011;43:1018–1021.
Beroukhim R, Getz G, Nghiemphu L et al. Assessing the significance of chromosomal aberrations in cancer: methodology and application to glioma. Proc Natl Acad Sci USA 2007;104:20007–20012.
Beroukhim R, Mermel CH, Porter D et al. The landscape of somatic copy-number alteration across human cancers. Nature 2010;463:899–905.
Mermel CH, Schumacher SE, Hill B et al. GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol 2011;12:R41.
Geigl JB, Speicher MR . Single-cell isolation from cell suspensions and whole genome amplification from single cells to provide templates for CGH analysis. Nat Protoc 2007;2:3173–3184.
Cheng DT, Mitchell TN, Zehir A et al. Memorial Sloan Kettering-Integrated Mutation Profiling of Actionable Cancer Targets (MSK-IMPACT): a hybridization capture-based next-generation sequencing clinical assay for solid tumor molecular oncology. J Mol Diagn 2015;17:251–264.
Krueger F, Trim Galore: a wrapper script to automate quality and adapter trimming as well as quality control. Available at http://www.bioinformatics.babraham.ac.uk/projects/trim_galore/ 2013 (accessed on 30 August 30).
Li H, Durbin R . Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009;25:1754–1760.
McKenna A, Hanna M, Banks E et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 2010;20:1297–1303.
Cibulskis K, Lawrence MS, Carter SL et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat Biotechnol 2013;31:213–219.
Rausch T, Zichner T, Schlattl A et al. DELLY: structural variant discovery by integrated paired-end and split-read analysis. Bioinformatics 2012;28:i333–i339.
van de Nes J, Gessi M, Sucker A et al. Targeted next generation sequencing reveals unique mutation profile of primary melanocytic tumors of the central nervous system. J Neurooncol 2016;127:435–444.
Lee CS, Bhaduri A, Mah A et al. Recurrent point mutations in the kinetochore gene KNSTRN in cutaneous squamous cell carcinoma. Nat Genet 2014;46:1060–1062.
Brash DE, Rudolph JA, Simon JA et al. A role for sunlight in skin cancer: UV-induced p53 mutations in squamous cell carcinoma. Proc Natl Acad Sci USA 1991;88:10124–10128.
Ateenyi-Agaba C, Dai M, Le Calvez F et al. TP53 mutations in squamous-cell carcinomas of the conjunctiva: evidence for UV-induced mutagenesis. Mutagenesis 2004;19:399–401.
Ziegler A, Leffell DJ, Kunala S et al. Mutation hotspots due to sunlight in the p53 gene of nonmelanoma skin cancers. Proc Natl Acad Sci USA 1993;90:4216–4220.
Horn S, Figl A, Rachakonda PS et al. TERT promoter mutations in familial and sporadic melanoma. Science 2013;339:959–961.
Huang FW, Hodis E, Xu MJ et al. Highly recurrent TERT promoter mutations in human melanoma. Science 2013;339:957–959.
Killela PJ, Reitman ZJ, Jiao Y et al. TERT promoter mutations occur frequently in gliomas and a subset of tumors derived from cells with low rates of self-renewal. Proc Natl Acad Sci USA 2013;110:6021–6026.
Morris LG, Kaufman AM, Gong Y et al. Recurrent somatic mutation of FAT1 in multiple human cancers leads to aberrant Wnt activation. Nat Genet 2013;45:253–261.
Nowell CS, Radtke F . Notch as a tumour suppressor. Nat Rev Cancer 2017;17:145–159.
Ranganathan P, Weaver KL, Capobianco AJ . Notch signalling in solid tumours: a little bit of everything but not all the time. Nat Rev Cancer 2011;11:338–351.
Rizzo P, Osipo C, Foreman K et al. Rational targeting of Notch signaling in cancer. Oncogene 2008;27:5124–5131.
Sanchez-Martin M, Ferrando A . The NOTCH1-MYC highway toward T-cell acute lymphoblastic leukemia. Blood 2017;129:1124–1133.
Agrawal N, Frederick MJ, Pickering CR et al. Exome sequencing of head and neck squamous cell carcinoma reveals inactivating mutations in NOTCH1. Science 2011;333:1154–1157.
South AP, Purdie KJ, Watt SA et al. NOTCH1 mutations occur early during cutaneous squamous cell carcinogenesis. J Invest Dermatol 2014;134:2630–2638.
Wang NJ, Sanborn Z, Arnett KL et al. Loss-of-function mutations in Notch receptors in cutaneous and lung squamous cell carcinoma. Proc Natl Acad Sci USA 2011;108:17761–17766.
Kamb A, Gruis NA, Weaver-Feldhaus J et al. A cell cycle regulator potentially involved in genesis of many tumor types. Science 1994;264:436–440.
Pleasance ED, Cheetham RK, Stephens PJ et al. A comprehensive catalogue of somatic mutations from a human cancer genome. Nature 2010;463:191–196.
Li YY, Hanna GJ, Laga AC et al. Genomic analysis of metastatic cutaneous squamous cell carcinoma. Clin Cancer Res 2015;21:1447–1456.
Pickering CR, Zhou JH, Lee JJ et al. Mutational landscape of aggressive cutaneous squamous cell carcinoma. Clin Cancer Res 2014;20:6582–6592.
Snyder A, Makarov V, Merghoub T et al. Genetic basis for clinical response to CTLA-4 blockade in melanoma. N Engl J Med 2014;371:2189–2199.
Rizvi NA, Hellmann MD, Snyder A et al. Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 2015;348:124–128.
Van Allen EM, Miao D, Schilling B et al. Genomic correlates of response to CTLA-4 blockade in metastatic melanoma. Science 2015;350:207–211.
Le DT, Uram JN, Wang H et al. PD-1 blockade in tumors with mismatch-repair deficiency. N Engl J Med 2015;372:2509–2520.
Acknowledgements
We thank Nadine Stadler and Iris Moll for excellent technical assistance. The presented study was supported by a grant from the Mercur-Stiftung. This research was funded in part through the NIH/NCI Cancer Center Support Grant P30 CA008748. The funders were not involved in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Modern Pathology website
Rights and permissions
About this article
Cite this article
Griewank, K., Wiesner, T., Murali, R. et al. Atypical fibroxanthoma and pleomorphic dermal sarcoma harbor frequent NOTCH1/2 and FAT1 mutations and similar DNA copy number alteration profiles. Mod Pathol 31, 418–428 (2018). https://doi.org/10.1038/modpathol.2017.146
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2017.146
This article is cited by
-
Atypical Fibroxanthoma Treated with a Topical Combination of Imiquimod, Tazarotene, and 5-Fluorouracil
Dermatology and Therapy (2024)
-
Systematic Review and Case Series of Pleomorphic Dermal Sarcoma: an Uncommon Cutaneous Pathology Predominantly in the Head and Neck
SN Comprehensive Clinical Medicine (2022)
-
The clinicopathologic spectrum and genomic landscape of de-/trans-differentiated melanoma
Modern Pathology (2021)
-
Soft Tissue Special Issue: Cutaneous Pleomorphic Spindle Cell Tumors
Head and Neck Pathology (2020)
-
Liquid Biopsy in der Tumordiagnostik
Wiener klinisches Magazin (2020)