Abstract
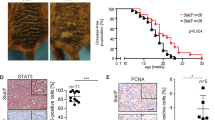
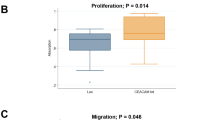
A large-scale gene expression study of melanoma metastases was performed to identify genes involved in late-stage tumor progression. Overall 248 genes, out of more than 47,000 tested, are differentially expressed when comparing peripheral areas (invasion front) with central tumor areas of melanoma metastases. As determined by gene ontology analysis, members of the STAT signaling pathway show significant enrichment. In particular, Stat1 is highly expressed in peripheral compared with central tumor areas. In line with this, stable knockdown of STAT1 in metastatic melanoma cells significantly impairs their migratory and invasive capacity in wounding and matrigel assays. Moreover, STAT1 knockdown affects the metastatic behavior of melanoma cells in a mouse model of melanoma metastasis. Taken together, these data suggest that Stat1 might play a role in late-stage melanoma progression. Interference with the Stat1 pathway could have therapeutic implications for late-stage melanoma patients.



Similar content being viewed by others
References
Miller AJ, MCJr Mihm (2006) Melanoma. N Engl J Med 355(1):51–65
Haass NK, Smalley KSM, Li L, Herlyn M (2005) Adhesion, migration and communication in melanocytes and melanoma. Pigment Cell Res 18(3):150–159
Greene VR, Johnson MM, Grimm EA et al (2009) Frequencies of NRAS and BRAF mutations increase from the radial to the vertical growth phase in cutaneous melanoma. J Invest Dermatol 129(6):1483–1488
Norton L, Massagué J (2006) Is cancer a disease of self-seeding? Nat Med 12(8):875–878
Brierley MM, Fish EN (2005) Stats: multifaceted regulators of transcription. J Interferon Cytokine Res 25(12):733–744
Levy DE, Darnell JE Jr (2002) Stats: transcriptional control and biological impact. Nat Rev Mol Cell Biol 3(9):651–662
Shuai K, Schindler C, Prezioso VR et al (1992) Activation of transcription by IFN-gamma: tyrosine phosphorylation of a 91-kD DNA binding protein. Science 258(5098):1808–1812
Schindler C, Levy DE, Decker T (2007) JAK-STAT signaling: from interferons to cytokines. J Biol Chem 282(28):20059–20063
Chapgier A, Boisson-Dupuis S, Jouanguy E et al (2006) Novel STAT1 alleles in otherwise healthy patients with mycobacterial disease. PLoS Genet 2(8):e131
Jaeger J, Koczan D, Thiesen HJ et al (2007) Gene expression signatures for tumor progression, tumor subtype, and tumor thickness in laser-microdissected melanoma tissues. Clin Cancer Res 13(3):806–815
Carlotti F, Bazuine M, Kekarainen T et al (2004) Lentiviral vectors efficiently transduce quiescent mature 3T3–L1 adipocytes. Mol Ther 9(2):209–217
Buettner R, Mora LB, Jove R (2002) Activated STAT signaling in human tumors provides novel molecular targets for therapeutic intervention. Clin Cancer Res 8(4):945–954
Hou SX, Zheng Z, Chen X, Perrimon N (2002) The JAK/STAT Pathway in model organisms: emerging roles in cell movement. Dev Cell 3(6):765–778
Silver DL, Montell DJ (2001) Paracrine signaling through the JAK/STAT pathway activates invasive behavior of ovarian epithelial cells in Drosophila. Cell 107(7):831–841
Silver DL, Naora H, Liu J et al (2004) Activated signal transducer and activator of transcription (STAT) 3: localization in focal adhesions and function in ovarian cancer cell motility. Cancer Res 64(10):3550–3558
Xie B, Zhao J, Kitagawa M et al (2001) Focal adhesion kinase activates STAT1 in integrin-mediated cell migration and adhesion. J Biol Chem 276(22):19512–19523
Bienz M, Clevers H (2003) Armadillo/β-catenin signals in the nucleus-proof beyond a reasonable doubt? Nat Cell Biol 5(3):179–182
Gütgemann A, Golob M, Müller S et al (2001) Isolation of invasion-associated cDNAs in melanoma. Arch Dermatol Res 293(6):283–290
Khodarev NN, Roach P, Pitroda SP et al (2009) STAT1 pathway mediates amplification of metastatic potential and resistance to therapy. PloS ONE 4(6):e5821
Khodarev NN, Minn AJ, Efimova EV et al (2007) Signal transducer and activator of transcription 1 regulates both cytotoxic and prosurvival functions in tumor cells. Cancer Res 67(19):9214–9220
Weichselbaum RR, Ishwaran H, Yoon T et al (2008) An interferon-related gene signature for DNA damage resistance is a predictive marker for chemotherapy and radiation for breast cancer. Proc Natl Acad Sci USA 105(47):18490–18495
Gupta GP, Massagué J (2006) Cancer metastasis: building a framework. Cell 127(4):679–695
Battle TE, Frank DA (2002) The role of STATs in apoptosis. Curr Mol Med 2(4):381–392
Hartman SE, Bertone P, Nath AK et al (2005) Global changes in STAT target selection and transcription regulation upon interferon treatments. Genes Dev 19(24):2953–2968
Wormald S, Hilton DJ, Smyth GK et al (2006) Proximal genomic localization of STAT1 binding and regulated transcriptional activity. BMC Genomics 7:254
Gatenby RA, Gillies RJ (2008) A microenvironmental model of carcinogenesis. Nat Rev Cancer 8(1):56–61
Acknowledgment
We thank R. Waterstradt and N. Harmel for excellent technical assistance. This study was supported in part by the Erich and Gertrud Roggenbuck Stiftung, Hamburg, Germany.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
10585_2010_9310_MOESM3_ESM.tif
Supplementary Fig. 1: Differential expression of Stat1 and Stat3 in melanoma metastases. Stat1 and Stat3 protein expression was analysed in 10 melanoma metastases by immunohistochemistry. Immunohistochemical staining of three different metastases (Meta1-3) is shown. Magnification x100. (TIFF 28835 kb)
10585_2010_9310_MOESM4_ESM.tif
Supplementary Fig. 2: Differential expression of STAT1 in knockdown and control cells in vivo. Real-time RT-PCR analysis of STAT1 expression in lung and gastrointestinal melanoma metastases of athymic nude mice intravenously injected with control (siCo) and STAT1 knockdown (siSTAT1) melanoma cells. (TIFF 1530 kb)
Rights and permissions
About this article
Cite this article
Schultz, J., Koczan, D., Schmitz, U. et al. Tumor-promoting role of signal transducer and activator of transcription (Stat)1 in late-stage melanoma growth. Clin Exp Metastasis 27, 133–140 (2010). https://doi.org/10.1007/s10585-010-9310-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10585-010-9310-7