Abstract
Huntington's disease results from a CAG repeat expansion within the Huntingtin gene; this is measured routinely in diagnostic laboratories. The European Huntington's Disease Network REGISTRY project centrally measures CAG repeat lengths on fresh samples; these were compared with the original results from 121 laboratories across 15 countries. We report on 1326 duplicate results; a discrepancy in reporting the upper allele occurred in 51% of cases, this reduced to 13.3% and 9.7% when we applied acceptable measurement errors proposed by the American College of Medical Genetics and the Draft European Best Practice Guidelines, respectively. Duplicate results were available for 1250 lower alleles; discrepancies occurred in 40% of cases. Clinically significant discrepancies occurred in 4.0% of cases with a potential unexplained misdiagnosis rate of 0.3%. There was considerable variation in the discrepancy rate among 10 of the countries participating in this study. Out of 1326 samples, 348 were re-analysed by an accredited diagnostic laboratory, based in Germany, with concordance rates of 93% and 94% for the upper and lower alleles, respectively. This became 100% if the acceptable measurement errors were applied. The central laboratory correctly reported allele sizes for six standard reference samples, blind to the known result. Our study differs from external quality assessment (EQA) schemes in that these are duplicate results obtained from a large sample of patients across the whole diagnostic range. We strongly recommend that laboratories state an error rate for their measurement on the report, participate in EQA schemes and use reference materials regularly to adjust their own internal standards.
Similar content being viewed by others
INTRODUCTION
Huntington's disease (HD) is an autosomal dominant progressive neurodegenerative disorder with onset usually, but not exclusively, in adult life. In 1993, the causative mutation was identified as an unstable CAG repeat expansion in the first exon of the Huntingtin (HTT) gene.1 This was rapidly introduced into clinical practice for diagnostic and predictive testing.2 Guidelines for reporting genetic test results were established by the American College of Medical Genetics (ACMG) and the American Society of Human Genetics, which state: a CAG repeat length of less than 27 is unequivocally normal; a CAG repeat length of 27–35 is normal but there may be expansion into the pathological range in future generations; a CAG repeat length of 36–39 is an abnormal result but there may be reduced penetrance; a CAG repeat length of 40 or more is unequivocally abnormal.3
External quality assessment (EQA) schemes have developed to ensure the accuracy of laboratory reporting.4 The model for these schemes is that validated aliquots of DNA samples from one or more patients with known genotypes are circulated to participating laboratories for analysis of the CAG repeat length; the reports generated are collated centrally and ideally, all participating laboratories should generate the same genotype results and reports with similar interpretations.
In this paper we present the results from a different model of assessing the accuracy of genotype reporting: a fresh blood sample was taken from a large sample of individuals participating in the European Huntington's Disease Network (EHDN) REGISTRY project; the DNA was extracted and CAG repeat length for both alleles was measured by a central laboratory and the result compared with that obtained from the local service laboratory.
MATERIALS AND METHODS
Patient Samples
The EHDN REGISTRY project5 is a multinational observational study; more details are available at http://www.euro-hd.net/html/registry. The data recorded from participants includes the result of the CAG repeat length reported by the local service laboratory. Participants have an option of donating fresh blood samples, which are taken in acid citrate dextrose (ACD tubes Vacutainer, Becton Dickinson, Milan, Italy) and couriered to the central laboratory, BioRep, Milan.
Laboratory Analysis
DNA was extracted from whole blood using the salting-out procedure,6 and the HTT gene CAG repeat length analysed by PCR amplification followed by capillary electrophoresis using the MegaBace Fragment Profiler Software from General Electric (Buckinghamshire, UK)7, 8 using the following primers:
HD1: 5′-FAM-ccttcgagtccctcaagtccttc-3′
HD5: 5′-cggctgaggcagcagcggctgt-3′
Hu4: 5′-HEX-atggcgaccctggaaaagctgatgaa-3′
Hu5: 5′-ggcggtggcggctgttgctgctgctgctgc-3′
Each sample was amplified with two primer sets: Hu4/Hu5, which amplifies CAG repeat sequences only and not adjacent CCG polymorphic variant. CCG and CAG are amplified with primer set HD1/HD5, this allows the detection of heterozygous CAG/CCG repeats. The reported results gave the CAG repeat number only.
BioRep participated in the EQA schemes organised by the European Molecular Genetics Quality Network (EMQN) on alternate years and the results are always congruent.
A second, independent, accredited laboratory in Tübingen, Germany, duplicated CAG repeat analyses for a subset of 348 DNA samples using the same DNA extracted at BioRep and the same primer sets. The amplified DNA was separated using the Applied Biosystems (Carlsbad, CA, USA) capillary electrophoresis. From 2002, the Tübingen laboratory has participated in the national (BVDH; 2002–2007) and European (EMQN, starting 2008) quality measures. All quality measure samples have been genotyped without any results outside error limits (accuracy within ±1 CAG for repeats below 42 CAG repeats; accuracy within ±1 for large HD mutations). The Tübingen lab has had a consistent discrepancy of −1 CAG allele in the lower repeat (which has been accounted for and corrected in this data series). The results obtained from the two laboratories were compared: the Tübingen values were subtracted from those of BioRep's.
In total, 1326 fresh samples collected between January 2004 and June 2009 were available for analysis. Duplicate results were compared by subtracting the BioRep result from that obtained from the local service laboratory for both the upper and lower allele. The number of samples reported from individual laboratories varied widely; hence, we aggregated the results for 10 countries contributing more than 20 samples to the study. We then applied the acceptable measurement errors to the discrepancies as proposed by the ACMG:9 these are: ±1 for CAG repeat lengths ≤43; ±2 for CAG repeat lengths between 44 and 50; ±3 for CAG repeat lengths between 51 and 75; ±4 for CAG repeat lengths >75.
The European Best Practice Guidelines (BPG) for HD is still in draft form;11 however, the suggested measurement errors in that document are: ±1 for CAG repeat lengths ≤42 and ±3 for CAG repeat lengths ≥43. We tested the effect of this standard on our data.
Use of Reference Materials
The US National Institute for Standards and Technology (NIST) developed standard reference materials for HD.10 The standards consisted of CAG repeat lengths of: sample 1, 15 and 29; sample 2, 7 and 36; sample 3, 15 and 40; sample 4, 35 and 45; sample 5, 39 and 50 and sample 6, 17 and 75. BioRep analysed these samples blind to the known CAG repeat size.
Participation in EMQN EQA
The names of laboratories participating in the 2009 EMQN EQA could not be disclosed for reasons of anonymity; however, the list of 121 laboratories participating in this study was sent to the EMQN coordinator who reported on the number participating in the 2009 EQA.
RESULTS
Duplicate results for the upper allele were available for 1307 patients and 19 non-mutation carrying controls who had a family history of HD. There were 654 (49%) concordant results and 672 (51%) discrepancies, which are summarised in Table 1: 31% were discrepant by one CAG repeat, 12% were discrepant by two CAG repeats and 8% had a discrepancy of three or more CAG repeats. The discrepancies were in both directions with 370 (55%) showing an increase, and 302 (45%) showing a decrease in size when re-analysed by BioRep. The discrepancy rate fell to 13.3% when we applied the acceptable measurement errors proposed by the ACMG and 9.7% when we applied the proposed BPG measurement error. The local results came from 121 laboratories, of these, 45 (31%) participated in the 2009 EMQN EQA. It is not known how many laboratories participated in national quality assessment schemes. Discrepancies came from 86 laboratories (71.1%): of these, 49 laboratories (40.5%) had discrepancies outside the proposed ACMG measurement error and 41 (33.9%) were outside the proposed BPG measurement error.
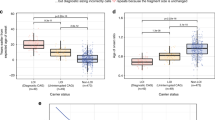
The number of samples each laboratory contributed to the study varied considerably; 103 laboratories contributed less than 20 local results, whereas 4 laboratories contributed 50 or more local results. We did not wish to identify individual laboratories and we did not think it was fair to compare the measurement error of a laboratory contributing more than 50 results with ones contributing only one or two results, hence, we aggregated the local laboratories’ results by country. The local laboratories were distributed across 15 countries; however, five each contributed less than 20 results to the study and were excluded from the analysis of discrepancy rate by country. Figure 1 shows the discrepancy rate for 10 countries, which represents 1276 duplicate samples (96.2% of the total) together with the effect of applying both the proposed ACMG- and Draft BPG-acceptable measurement errors. The results for country 1 suggest there is a consistent error in reporting the upper allele. There was a wide variation in the discrepancy rate by country even after allowing for acceptable measurement errors.
A discrepancy is clinically significant if it crosses a boundary at 35–36 or 39–40 CAG repeat lengths; this occurred in 52 (4%) patients. Results for the upper allele changed from the reduced to the full penetrance range in 36 cases, whereas in 11 cases they moved from the full to the reduced penetrance range. A potential misdiagnosis occurred in five (0.4%) cases, which changed at the critical 35–36 CAG repeat length boundary. In one case, this was because of a labelling error on the fresh sample sent to BioRep; this leaves four cases (0.3%) with an unexplained potential misdiagnosis.
Methods of measuring the CAG repeat length in local laboratories have developed since 1993; hence, we considered the effect of this on our data. Of the 672 discrepant samples, the year in which the local laboratory reported the result was known for 663/672 samples; 89 discrepancies occurred in the years 1993–1996, and 549 occurred in the period 2006–2009; these represent 66.3 and 46.6% of the total samples received for the respective time periods. Applying acceptable measurement errors from the ACMG for these two time periods gave discrepancy rates of 13.5% and 12.8%, respectively. Applying the Draft-proposed BPG measurement errors for the same periods gave discrepancy rates of 7.9% and 8.9%, respectively.
Duplicate results were available for the lower allele in 1250 cases, of which 757 (60.6%) were concordant. The 493 (39.4%) discordant results are summarised in Table 2. Following re-analysis by BioRep, 153 (31%) of the discrepancies were an increase in the CAG repeat size, and 340 (69%) were a decrease in CAG repeat size. Applying either the ACMG or the BPG measurement limits resulted in 160 (12.8%) cases being discordant. The two cases with a lower allele CAG repeat length of more than 36 were reported as having a result of 37 and 50 CAG repeats and 39 and 45 CAG repeats for the lower and upper allele, respectively, by both the local service laboratory and BioRep.
In the case of a discordant result, it is not possible to say whether the local laboratory or BioRep gave the correct CAG repeat size. We tested the reliability of BioRep in two ways. First, a subset of 348 DNA samples was analysed at a second laboratory in Tübingen. The results were concordant for the upper and lower alleles in 324 (93%) and 327 (94%) cases, respectively. The upper allele decreased in size by 1 CAG repeat in 17 cases and increased by 1 CAG repeat in 7 cases at Tübingen. For the lower allele, 17 cases decreased and 4 cases increased in size by 1 CAG repeat at Tübingen. This represents complete concordance if either the ACMG- or BPG-proposed measurement errors are applied. In three cases, the discrepancy of 1 CAG repeat occurred in both the upper and lower allele. Second, BioRep was unaware of the known CAG repeat length for the six reference samples supplied by NIST; 11 alleles were reported correctly, but the 75 CAG standard was reported as 74, which is well within the margin of error for this allele size.
DISCUSSION
In a series of 1326 duplicate samples, we found a discrepancy in reporting the upper CAG repeat allele outside the ACMG- and BPG-proposed measurement errors in 13.3% and 9.7% of cases, respectively. The observation that the discrepancies were in both directions suggests that the result was not due to a consistent error at BioRep.
We considered the possibility that the explanation for our results is that the discrepancies occurred in earlier years; however, 41% of the samples in this study came from the period 2006–2009, with discrepancy rates outside the proposed ACMG and BPG acceptable measurement errors of 12.8% and 8.9%, respectively, indicating that there is a current problem, which needs to be both recognised and addressed.
The purpose of this report is not to identify any specific laboratory, but rather to address the question of whether the problem is widespread or confined to just a few laboratories. The observation that the number of laboratories with discrepancies outside the proposed ACMG- and BPG-acceptable measurement errors is 49 and 41, respectively; in addition, the data from Figure 1 indicates that the problem occurs across Europe and is not confined to few countries. Country 1 had a significant number of samples discrepant by 1 CAG repeat, which is within acceptable measurement limits. The difference in measurement between two laboratories does not allow us to say which is the correct measurement, but the issue may well be resolved by more widespread use of standard reference materials.
There have been previous reports of discrepancies in the measurement of CAG repeat length. The 1999 report of the EQA scheme for HD, organised by the EMQN, asked laboratories to report on five samples with set measurement limits of ±1 repeat for values up to 40, and ±3 repeats for results of more than 40 repeats; 6.2% of results fell outside these limits and there was a potential misdiagnosis rate of 1.3%.4 Although the methods adopted by an EQA and our study are not directly comparable, we note that follow-up data reported in 2009, show that after more than 10 years of EQA, based on three samples being circulated among laboratories, a potential misdiagnosis rate of 1–4% still occurred.12
In a study of reduced penetrance alleles, 200 samples were collected from centres reporting results in this range, but 24 (12%) had to be excluded because duplicate results from two reference laboratories showed that these had results of ≥40 repeats.13 In the same study, a total of 238 duplicate results were available from the two reference centres; 10 (4.2%) differed by one CAG repeat in the reporting of the upper allele, and in 2 (0.84%) of these cases, the discrepancy was between 39 and 40 repeats.13
Our study was based on duplicate reporting of results for 1307 patients across the whole spectrum of CAG repeat length, and showed that discrepancies in measuring and reporting this are common; clinically significant discrepancies occurred in reporting the upper allele in 4.0% of the sample, with an unexplained diagnostic error rate in 0.3% in this large patient series. The fact that our study considered results from across the diagnostic range may explain why the unexplained misdiagnosis rate is less than that suggested from the EQA studies, which are more likely to concentrate on samples at clinically important boundaries.
As part of the standard operating procedure for our REGISTRY study, principal investigators receive a periodic printout of the results obtained from BioRep together with the patient's pseudonym so they are in a position to check a specific result. If a patient has a CAG repeat length well above 40 repeats and there is a discrepancy of one or more CAG repeats then this may not have any clinical consequence but, knowledge of the degree of discrepancy that can occur may be important for those undertaking research, based on CAG repeat length data pooled from multiple service laboratories.
We have neither identified the countries contributing samples nor have we sought to identify specific laboratories within countries; our aim is to draw attention to the wide variation, which exists across Europe and that it is not specific to one country. It is not possible to explain the cause of the variation in the results between laboratories from this study but one possibility could be the use of different electrophoretic matrices.14 The data do not allow us to investigate all the possible causes of the discrepancies but, having identified that a problem exists, we wish to suggest ways of addressing the issues. Our result suggests that laboratories should quote an error rate in reporting the allele sizes; this error rate may increase as the allele size increases. There are no universally agreed acceptable measurement errors hence we recommend laboratories determine the error rate experimentally rather than choosing rates from a published recommendation. This will allow clinicians to be aware that the result is not an absolute but, as with all measurements, is subject to error. Discrepancies outside acceptable measurement limits continue to occur, despite the participation of laboratories in EQA schemes; use of standard reference materials to calibrate internal standards should result in a further reduction of the scatter in measurements.
Cases of clear misdiagnosis may be rare, but there may be more problems at the 39/40 boundary; the genetic counselling differs slightly depending on whether the result is in the reduced penetrance or unequivocally abnormal range. We strongly recommend that service laboratories should participate in EQA schemes, but, in addition, should use known reference materials regularly to adjust their own internal standards.
References
Huntington's Disease Collaborative Research Group: A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington's disease chromosomes. Cell 1993; 72: 971–983.
Kremer B, Godberg P, Andrew SE et al: A worldwide study of the Huntington's disease mutation. The sensitivity and specificity of measuring CAG repeats. New Engl J Med 1994; 330: 1401–1406.
ACMG/ASHG Statement: Laboratory guidelines for Huntington's disease genetic testing. Am J Hum Genet 1998; 62: 1243–1247.
Losekoot M, Bakker B, Laccone F, Stenhouse S, Elles R : A European pilot quality assessment scheme for molecular diagnosis of Huntington's disease. Eur J Hum Genet 1999; 7: 217–222.
Orth M, Handley OJ, Schwenke C et al: Observing Huntington's disease: the European Huntington's Disease Network's REGISTRY [Internet]. Version 19. PLoS Curr: Huntington Dis 28 September 2010 (revised 13 April 2011): PMC2947793.
Miller SA, Dykes DD, Polesky HF : A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res 1988; 16: 1215.
Warner JP, Barron LH, Brock DJ : A new polymerase chain reaction (PCR) assay for the trinucleotide repeat that is unstable and expanded on Huntington's disease chromosomes. Mol Cell Probes 1993; 7: 235–239.
Riess O, Noerremoelle A, Soerensen SA, Epplen JT : Improved PCR conditions for the stretch of (CAG)n repeats causing Huntington′s disease. Hum Mol Genet 1993; 2: 637.
American College of Medical Genetics Standards and Guidelines for Clinical Genetics Laboratories, 2006 Edition. Available at: http://www.acmg.net/Pages/ACMG_Activities/stds-2002/HD.htm.
Kline M : Certification of Standard Reference Material for the CAG repeats of Huntington Disease, 2010, Available at: http://www.nist.gov/mml/biochemical/genetics/huntington_disease_srm.cfm.
Barton D : Best Practice Guidelines for the molecular analysis of Huntington's disease, 2010, (Personal communication). This draft guideline may be accessed at: http://www.cmgs.org/BPGs/pdfs%20current%20bpgs/HD%20BPG%20April%202010%20for%20consultation.pdf.
Losekoot M, Seneca S, Kamarainen O, Patton S : Ten years’ experience in external quality assessment for genetic testing of Huntington's disease in Europe: still room for improvement. Clin Genet 2009; 76 (Suppl 1): 104–105.
Quarrell OW, Rigby AS, Baron L et al: Reduced penetrance alleles for Huntington's disease: a multicentre study. J Med Genet 2007; 44: e68.
Williams LC, Hegde MR, Herrera G, Stapleton PM, Love DR : Comparative semi-automated analysis of (CAG) repeats in the Huntington disease gene: use of internal standards. Mol Cell Probes 1999; 13: 283–289.
Acknowledgements
The REGISTRY project is funded by the Cure Huntington's Disease Initiative. We thank Simon Patton from EMQN for reviewing the list of laboratories participating in this study and stating the number participating in the 2009 EMQN EQA.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Competing interests
G Bernhard Landwehrmeyer received research grants from CHDI and honoraria from Siena Biotech for consultations as well as compensation for the conduct of clinical trials from Amarin, Neurosearch, Novartis and Medivation. The remaining authors declare no conflict of interest.
Appendix
Appendix
Collaborators 2004–2009
Language coordinators
Katrin Barth, Leonor Correia Guedes, Ana Maria Finisterra, Monica Bascuñana Garde, R Bos, Daniel Ecker, Christine Held, Kerstin Koppers; Mathilde Laurà, Asunción Martínez Descals, Tim McLean, Tiago Mestre, Sara Minster, Daniela Monza, Jenny Townhill (formerly Naji), Michael Orth, Helene Padieu, Laurent Paterski; Nadia Peppa, Susana Pro Koivisto, Amandine Rialland, Ninni Røren (formerly Heinonen) Pavla Šašinková, Patricia Trigo Cubillo, Marlene R van Walsem, Marie-Noelle Witjes-Ané, Elizaveta Yudina (formerly Tarasova), Daniel Zielonka, Eugeniusz Zielonka; Paola Zinzi.
Registry Steering Committee
A-C Bachoud-Lévi, AR Bentivoglio, I Biunno, R Bonelli, J-M Burgunder, SB Dunnett, JJ Ferreira, OJ Handley, A Heiberg, T Illmann, GB Landwehrmeyer, J Levey, JE Nielsen, M Päivärinta, RAC Roos, A Rojo Sebastián, SJ Tabrizi, W Vandenberghe, C Verellen-Dumoulin, J Zaremba, T Uhrova, J Wahlström.
Austria
Graz (LKH Graz, Abteilung für Psychiatrie): Raphael M. Bonelli; Brigitte Herranhof; Anna Holl (formerly Hödl); Hans-Peter Kapfhammer; Michael Koppitz; Markus Magnet; Daniela Otti; Annamaria Painold; Karin Reisinger; Monika Scheibl; Karen Hecht; Sabine Lilek; Nicole Müller; Helmut Schöggl; Jasmin Ullah.
Innsbruck (Neurologie, Universitätsklinik Innsbruck): Florian Brugger; Caroline Hepperger; Anna Hotter; Philipp Mahlknecht; Michael Nocker; Klaus Seppi; Gregor Wenning; Lisa Buratti; Eva-Maria Hametner; Christiane Holas; Anna Hussl; Katharina Mair; Werner Poewe; Elisabeth Wolf; Anja Zangerl; Eva-Maria Braunwarth.
Belgium
Charleroi (Institut de Pathologie et de Génétique): Pascale Ribaï. Brussels (VUB Neurology): Anja Flamez; V Morez; Sylvie de Raedt.
Leuven: (Universitair Ziekenhuis Gasthuisberg,): Andrea Boogaerts; Wim Vandenberghe; Dimphna van Reijen.
Czech Republic
Prague (Extrapyramidové centrum, Neurologická klinika, 1. LF UK a VFN): Jiř Klempíř; Martin Kucharík; Jan Roth.
DENMARK
Copenhagen (Hukommelsesklinikken, Rigshospitalet; Panum Instituttet): Jørgen E. Nielsen, Lena E. Hjermind, Oda Jakobsen, Jette Stokholm; Lis Hasholt, Anne Nørremølle, Sven Asger Sørensen.
Finland
Turku-Suvituuli (Rehabilitation Centre Suvituuli): Heli Hiivola; Kirsti Martikainen; Katri Tuuha.
Helsinki-The Family Federation of Finland (Department of Medical Genetics): Maarit Peippo; Marjatta Sipponen.
Germany
Aachen (Universitätsklinikum Aachen, Neurologische Klinik): Christoph Michael Kosinski; Eva Milkereit; Daniela Probst; Christian Sass; Johannes Schiefer; Christiane Schlangen; Cornelius J Werner.
Berlin (Klinik und Poliklinik für Neurologie – Charité – Universitätsmedizin Berlin): Harald Gelderblom; Josef Priller; Harald Prüß; Eike Jakob Spruth.
Bochum (Huntington-Zentrum (NRW) Bochum im St Josef-Hospital): Jürgen Andrich; Rainer Hoffmann; Peter H Kraus; Sabine Muth; Christian Prehn, Carsten Saft; Stephan Salmen; Christiane Stamm; Tanja Steiner; Katrin Straßburger.
Dinslaken (Reha Zentrum in Dinslaken im Gesundheitszentrums Lang): Herwig Lange.
Dresden (Universitätsklinikum Carl Gustav Carus an der Technischen Universität Dresden, Klinik und Poliklinik für Neurologie): Andrea Friedrich; Ulrike Hunger; Matthias Löhle; Simone Schmidt; Alexander Storch; Annett Wolz; Martin Wolz.
Freiburg (Neurologie, Universitätsklinik Freiburg): Johann Lambeck, Birgit Zucker.
Hamburg (Universitätsklinikum Hamburg-Eppendorf, Klinik und Poliklinik für Neurologie): Kai Boelmans; Christos Ganos; Ute Hidding; Jan Lewerenz; Alexander Münchau; Michael Orth; Jenny Schmalfeld; Lars Stubbe; Simone Zittel.
Heiligenhafen (Psychatrium Heiligenhafen): Walburgis Heinicke.
Marburg KPP (Klinik für Psychiatrie und Psychotherapie Marburg-Süd): Bernhard Longinus.
Marburg Uni (Neurologie, Universität Marburg): Kathrin Bürk; Jens Carsten Möller; Ida Rissling.
München (Huntington-Ambulanz im Neuro-Kopfzentrum – Klinikum rechts der Isar der Neurologischen Klinik und Poliklinik der Technischen Universität München): Alexander Peinemann; Michael Städtler; Adolf Weindl.
Münster (Universitätsklinikum Münster, Klinik und Poliklinik für Neurologie): Natalie Bechtel; Heike Beckmann; Stefan Bohlen; Eva Hölzner; Herwig Lange; Ralf Reilmann; Stefanie Rohm, Silke Rumpf ; Sigrun Schepers.
Taufkirchen (Isar-Amper-Klinikum – Klinik Taufkirchen (Vils)): Antonie Beister; Matthias Dose; Kathrin Hammer; Janina Kieni; Gabriele Leythaeuser; Ralf Marquard; Tina Raab; Sven Richter; Amina Selimbegovic-Turkovic; Caroline Schrenk; Michele Schuierer; Alexandra Wiedemann.
Ulm (Neurologie, Universitätsklinikum Ulm): Katrin Barth; Andrea Buck; Julia Connemann; Daniel Ecker; Carolin Eschenbach; Christine Held; Bernhard Landwehrmeyer; Franziska Lezius; Solveig Nepper; Anke Niess; Michael Orth, Sigurd Süßmuth, Sonja Trautmann; Patrick Weydt.
Italy
Bari (Dipartimento di Scienze Neurologiche e Psichiatriche Universita′ di Bari): Claudia Cormio; Olimpia Difruscolo; Vittorio Sciruicchio; Claudia Serpino; Marina de Tommaso.
Bologna (Dipartimento di Scienze Neurologiche – Clinica Neurologica, Università di Bologna): Sabrina Capellari; Pietro Cortelli; Roberto Gallassi; Roberto Poda; Giovanni Rizzo; Cesa Scaglione.
Florence (Dipartimento di Scienze Neurologiche e Psichiatriche Universita’ degli Studi di Firenze-Azienda Ospedaliera Universitaria Careggi): Elisabetta Bertini; Elena Ghelli, Andrea Ginestroni, Francesca Massaro, Claudia Mechi; Marco Paganini; Silvia Piacentini; Silvia Pradella; Anna Maria Romoli; Sandro Sorbi.
Genoa (Dipartimento di Neuroscienze, Oftalmologia e Genetica. Università di Genova): Giovanni Abbruzzese; Monica Bandettini di Poggio; Emilio Di Maria, Giovanna Ferrandes; Paola Mandich; Roberta Marchese.
Milan (Fondazione IRCCS Istituto Neurologico Carlo Besta): Alberto Albanese; Daniela Di Bella; Stefano Di Donato; Cinzia Gellera; Silvia Genitrini; Caterina Mariotti; Daniela Monza; Lorenzo Nanetti; Dominga Paridi; Paola Soliveri; Chiara Tomasello.
Naples (Dipartimento di Scienze Neurologiche Azienda Ospedaliera Universitaria Federico II): Giuseppe De Michele; Luigi Di Maio; Carlo Rinaldi; Cinzia Valeria Russo, Elena Salvatore; Tecla Tucci.
Pozzilli (IS) (Unità Operativa di Neurogenetica – IRCCS Neuromed): Milena Cannella; Valentina Codella, Francesca De Gregorio, Nunzia De Nicola, Tiziana Martino; Maria Simonelli; Ferdinando Squitieri.
Rome (Istituto di Neurobiologia e Medicina Molecolare & Istituto di Scienze e Tecnologie della Cognizione/CNR; Istituto di Neurologia Università Cattolica del Sacro Cuore): Anna Rita Bentivoglio; Claudio Catalli; Raffaella Di Giacopo; Alfonso Fasano; Marina Frontali; Arianna Guidubaldi; Tamara Ialongo; Gioia Jacopini; Giovanna Loria; Anna Modoni; Carla Piano; Piccininni Chiara; Davide Quaranta; Silvia Romano; Francesco Soleti; Maria Spadaro; Paola Zinzi.
The Netherlands
Enschede (Medisch Spectrum Twente): Monique SE van Hout; Jeroen PP van Vugt; A Marit de Weert.
Groningen (Polikliniek Neurologie): JJW Bolwijn; M Dekker; KL Leenders; JCH van Oostrom.
Leiden (Leiden University Medical Centre): Reineke Bos; Eve M. Dumas; Caroline K Jurgens; Simon JA van den Bogaard; Raymund AC Roos; Ellen P ‘t Hart; Marie-Noëlle Witjes-Ané.
Nijmegen (Department of Neurology, Universitair Medisch Centrum St Radboud): Berry Kremer; CCP Verstappen.
Norway
Oslo University Hospital (Rikshospitalet, Department of Medical Genetics and Department of Neurology): Arvid Heiberg; Marleen R van Walsem; Olaf Aaserud; Raghild Wehus.
Kathrine Bjørgo; Madelein Fannemel; Per Gørvell; Susana Pro Koivisto; Lars Retterstøl; Torborg Overland; Bodil Stokke.
Trondheim (St Olavs Hospital): Inga Bjørnevoll; Sigrid Botne Sando.
Poland
Gdansk (Specialist Hospital, Gdansk Zaspa): Emilia Sitek; Jaroslaw Slawek; Witold Soltan.
Katowice (Silesian Medical University, Katowice): Magdalena Boczarska-Jedynak; Barbara Jasinska-Myga; Gregorz Opala; Gabriela Kłodowska – Duda.
Krakow (Krakowska Akademia Neurologii): Krzysztof Banaszkiewicz; Andrzej Szczudlik; Monika Rudziñska; Magdalena Wójcik; Małgorzata Dec; Malgorzata Krawczyk.
Poznan (Medical University of Poznañ): Anna Bryl; Anna Ciesielska; Aneta Klimberg; Jerzy Marcinkowski; Justyna Sempołowicz; Daniel Zielonka; Husam Samara.
Warsaw-MU (Department of Neurology, Medical University of Warsaw): Piotr Janik; Anna Kalbarczyk; Hubert Kwiecinski; Zygmunt Jamrozik.
Warsaw-IPiN (Department of Genetics, Department of Neurology, Institute of Psychiatry and Neurology): Jakub Antczak; Katarzyna Jachinska; Maria Rakowicz; Przemyslaw Richter; Danuta Ryglewicz; Grzegorz Witkowski; Elzbieta Zdzienicka; Jacek Zaremba; Anna Sułek, Wioletta Krysa.
Portugal
Lisbon – (Hospital de Santa Maria; Neurological Clinical Research Unit, Instituto de Medicina Molecular): Tiago Mestre; Leonor Guedes; Miguel Coelho; Tiago Mendes; Anabela Valadas; Joaquim J Ferreira.
Porto – (Hospital Geral de Santo António): Sara Cavaco; Joana Damásio; Marina Magalhães.
Porto – (Hospital de São João): Miguel Gago; Carolina Garrett; Maria Rosália Guerra.
Spain
Granada (Neurología, Hospital Universitario San Cecilio de Granada): Francisco Barrero; Blas Morales.
Burgos (Servicio de Neurología Hospital General Yagüe): Esther Cubo; Natividad Mariscal; Jesús Sánchez.
Fuenlabrada (Hospital Universitario de Fuenlabrada): Fernando Alonso-Frech; Maria Rabasa Perez.
Madrid-Clinico (Hospital Clínico Universitario San Carlos): María Fenollar; Rocío García-Ramos García; Purificacion Pin Quiroga; Susana Vázquez Rivera; Clara Villanueva.
Madrid RYC (Hospital Ramón y Cajal, Neurología): Mónica Bascuñana; Marta Fatás Ventura; Guillermo García Ribas; Justo García de Yébenes; José Luis López – Sendón Moreno, Patricia Trigo Cubillo.
Madrid FJD (Madrid-Fundación Jiménez Díaz): Pedro J García Ruíz, Asunción Martínez-Descals, María José Saiz Artiga; Vicenta Sánchez.
Murcia (Hospital Universitario Virgen de la Arrixaca): María Fuensanta Noguera Perea; Fortuna Lorenza; María Martirio Antequera Torres; Gema Reinante ; Laura Vivancos Moreau.
Barcelona-Hospital Mútua de Terrassa : Miquel Aguilar Barbera; Dolors Badenes Guia; Laura Casas Hernanz ; Judit López Catena; Ana Rojo Sebastián, Pilar Quiléz Ferrer; Gemma Tome Carruesco.
Barcelona-Bellvitge (Hospital Universitari de Bellvitge): Jordi Bas; Núria Busquets Matilde Calopa.
Barcelona-Merced (Hospital Mare de Deu de La Merced): Marina Dalmau Elorza; Cristóbal Díez-Aja López; Santiago Durán-Sindreu Terol; Misericordia Floriach Robert; Belén Garzón Ruíz; Ana González Casado; Isabel Haro Martínez; Celia Mareca Viladrich; Regina Pons i Càrdenas; Elvira Roca; Joan Roig Llesoy; Jesús Miguel Ruiz Idiago; Mar Ruíz Vergara; Socorro Soriano García; Antonio Villa Riballo.
Palma (Hospital Son Dureta): Aranzazú Gorospe; Inés Legarda; Penelope Navas Arques; María José Torres Rodríguez, Barbara Vives.
Pamplona (Department of Medical Genetics, Hospital Virgen del Camino): Itziar Gaston.
Valencia (Hospital Universitari La Fe de Valencia): Maria Bosca; Juan Andres Burguera; Anabel Campos Garcia
Sweden
Stockholm (Karolinska University Hospital): Sven E Pålhagen; Martin Paucar; Per Svenningsson; Tina Walldén Reza-Soltani; Arja Höglund; Britta Sandström.
Göteborg (Sahlgrenska University Hospital): Jan Wahlström; Ulrika Høsterey-Ugander; Gunnel Fredlund; Radu Constantinescu; Liselotte Neleborn-Lingefjärd.
Stockholm-Ersta (NeuroHealth Consulting Sweden HB, Karolinska Institute): Joakim Tedroff ; Mona Esmaeilzadeh; Elisabeth Winnberg.
Switzerland
Bern: Jean-Marc Burgunder; Yvonne Burgunder; Yanik Stebler (Neurologische Klinik des Inselspitals); Alain Kaelin; Irene Romero; Michael Schüpbach; Sabine Weber Zaugg (Zentrum für Bewegungsstörungen, Neurologische Klinik und Poliklinik).
United Kingdom
Aberdeen (NHS Grampian, Clinical Genetics Centre): Roisin Jack; Kirsty Matheson; Zosia Miedzybrodzka; Daniela Rae; Sheila Simpson; Fiona Summers; Alexandra Ure.
Birmingham (The Barberry Centre, Department of Psychiatry): Jenny Crooks; Adrienne Curtis; Jenny de Souza (Keylock); Hugh Rickards; Jan Wright.
Cambridge (Cambridge Centre for Brain Repair, Forvie Site): Roger A. Barker; Anna Di Pietro; Kate Fisher; Anna Goodman; Susan Hill; Ann Kershaw; Sarah Mason; Nicole Paterson; Lucy Raymond.
Cardiff (The Institute of Medical Genetics, University Hospital of Wales): Jonathan Bisson, Monica Busse; Catherine Clenaghan; Lynda Ellison-Rose; Olivia Handley, Sarah Hunt, Jenny Townhill (formerly Naji); Kathleen Price; Anne Rosser.
Edinburgh (Molecular Medicine Centre, Western General Hospital, Department of Clinical Genetics): Maureen Edwards; Teresa Hughes (Scottish Huntington's Association); Marie McGill; Pauline Pearson; Mary Porteous; Paul Smith (Scottish Huntington's Association); Adam Zeman.
Exeter (Heavitree Hospital): Aaron Causley; Timothy Harrower, Debbie Howcroft; Nicol Lambord; Julia Rankin.
Fife (Scottish Huntington's Association Whyteman's Brae Hospital): Peter Brockie; Jillian Foster; Nicola Johns; Sue McKenzie, Jean Rothery, Gareth Thomas, Shona Yates.
Glasgow (Abercromby Centre): Joanne Miller; Stuart Ritchie.
Gloucester (Department of Neurology, Gloucestershire Royal Hospital): Liz Burrows; Amy Fletcher; Alison Harding, Fiona Laver; Mark Silva; Aileen Thomson.
Leeds (Department of Clinical Genetics, Chapel Allerton Hospital): Kathy Barnes; Carol Chu; Emma Hobson; Stuart Jamieson; Ivana Markova; Jenny Thomson; Jean Toscano; Sue Wild; Pam Yardumian.
Leicester (Leicestershire Partnership Trust, Mill Lodge): Colin Bourne; Carole Clayton; Heather Dipple; Jackie Clapton, Janet Grant; Diana Gross; Caroline Hallam; Julia Middleton; Ann Murch, Dawn Patino.
London (Guy's Hospital): Thomasin Andrews; Andrew Dougherty; Fred Kavalier; Charlotte Golding; Alison Lashwood; Dene Robertson; Deborah Ruddy; Anna Whaite.
London (St Georges-Hospital): Michael Patton, Maria Peterson; Sarah Rose.
London (The National Hospital for Neurology and Neurosurgery): Thomasin Andrews; Stefania Bruno; Elvina Chu; Karen Doherty; Charlotte Golding; Susie Henley; Nayana Lahiri; Marianne Novak; Aakta Patel; Joy Read; Elisabeth Rosser; Miranda Say; Sarah Tabrizi; Rachel Taylor; Thomas Warner; Edward Wild.
Manchester (Genetic Medicine, University of Manchester, Manchester Academic Health Sciences Centre and Central Manchester University Hospitals NHS Foundation Trust): Natalie Arran; Jenny Callaghan; David Craufurd; Ruth Fullam; Liz Howard; Susan Huson; Emma Oughton; Lucy Partington-Jones;); Julie Snowden; Andrea Sollom; Cheryl Stopford; Jennifer Thompson; Iris Trender-Gerhad; Nichola Verstraelen (formerly Ritchie); Leann Westmoreland.
Oxford (Oxford Radcliffe Hospitals NHS Trust): Andrea H Nemeth; Gill Suida.
Plymouth (Heathleigh Unit, Mount Gould Hospital): David Harrison; Max Hughes; Andrew Parkinson; Beverley Soltysiak.
Sheffield (The Royal Hallamshire Hospital – Sheffield Children's Hospital): Oliver Bandmann; Alyson Bradbury, Paul Gill, Helen Fairtlough, Kay Fillingham, Isabella Foustanos; Nadia Peppa, Katherine Tidswell.
Rights and permissions
About this article
Cite this article
Quarrell, O., Handley, O., O'Donovan, K. et al. Discrepancies in reporting the CAG repeat lengths for Huntington's disease. Eur J Hum Genet 20, 20–26 (2012). https://doi.org/10.1038/ejhg.2011.136
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2011.136
Keywords
This article is cited by
-
EMQN/CMGS best practice guidelines for the molecular genetic testing of Huntington disease
European Journal of Human Genetics (2013)