Abstract
Purpose
To assign functional properties to gene expression profiles of cervical cancer stages and identify clinically relevant biomarker genes.
Experimental design
Microarray samples of 24 normal and 102 cervical cancer biopsies from four publicly available studies were pooled and evaluated. High-quality microarrays were normalized using the CONOR package from the Bioconductor project. Gene expression profiling was performed using variance-component analysis for accessing most reliable probes, which were subsequently processed by gene set enrichment analysis.
Results
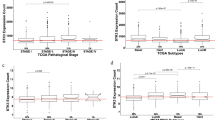
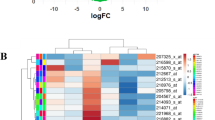
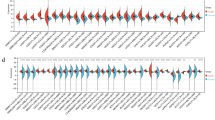
Of 22.277 probes that were subject to variance-component analysis, eleven probes had low heterogeneity, that is, a W/T ratio between 0.18 and 0.38. Seven of these probes are induced in all cervical cancer stages: they are GINS1, PAK2, DTL, AURKA, PRKDC, NEK2 and CEP55. The other four probes are induced in normal cervix: P11, EMP1, UPK1A and HSPC159. We performed GSEA of 9.873 probes exhibiting less variability, that is, having a W/T ratio of <0.75. Repeatedly, significant gene expression signatures were found that are related to treatment using angiocidin and darapladib. Additionally, expression signatures from immunological disease signatures were found, for example graft versus host disease and acute kidney rejection. Another finding comprises a gene expression signature in stage IB2 that refers to MT1-MMP-dependent migration and invasion. This gene signature is accompanied by gene expression signatures which refer to ECM receptor-mediated interactions.
Conclusion
Analysis of cervical cancer patient gene expression data reveals a novel perspective on HPV-mediated transcription processes. This novel point of view contains a better understanding and even might provide improvements to cancer therapy.




Similar content being viewed by others
References
Bachtiary B, Boutros PC, Pintilie M, Shi W, Bastianutto C, Li J-H, Schwock J, Zhang W, Penn LZ, Jurisica I, Fyles A, Liu F–F (2006) Gene expression profiling in cervical cancer: an exploration of intratumor heterogeneity. Clin Cancer Res 12:5632–5640. doi:10.1158/1078-0432.CCR-06-0357
Barrett T, Suzek TO, Troup DB et al (2005) NCBI GEO: mining millions of expression profiles–database and tools. Nucleic Acids Res 33:D562–D566. doi:10.1093/nar/gki022
Bidus MA, Risinger JI, Chandramouli GV, Dainty LA, Litzi TJ, Berchuck A, Barrett JC, Maxwell GL (2006) Prediction of lymph node metastasis in patients with endometrioid endometrial cancer using expression microarray. Clin Cancer Res 12(1):83–88. doi:10.1158/1078-0432.CCR-05-0835
Browne EP, Wing B, Coleman D, Shenk T (2001) Altered cellular mRNA levels in human cytomegalovirus-infected fibroblasts: viral block to the accumulation of antiviral mRNAs. J Virol 75(24):12319–12330. doi:10.1128/JVI.75.24.12319-12330.2001
Flechner SM, Kurian SM, Head SR, Sharp SM, Whisenant TC, Zhang J, Chismar JD, Horvath S, Mondala T, Gilmartin T, Cook DJ, Kay SA, Walker JR, Salomon DR (2004) Kidney transplant rejection and tissue injury by gene profiling of biopsies and peripheral blood lymphocytes. Am J Transplant 4:1475–1489. doi:10.1111/j.1600-6143.2004.00526.x
Gaurnier-Hausser A, Rothman VL, Dimitrov S, Tuszynski GP (2008) The novel angiogenic inhibitor, angiocidin, induces differentiation of monocytes to macrophages. Cancer Res 68(14):5905–5914. doi:10.1158/0008-5472.CAN-07-6179
Insinga A, Monestiroli S, Ronzoni S, Gelmetti V, Marchesi F, Viale A, Altucci L, Nervi C, Minucci S, Pelicci PG (2005) Inhibitors of histone deacetylases induce tumor-selective apoptosis through activation of the death receptor pathway. Nat Med 11:71–76. doi:10.1038/nm1160
Irizarry R, Hobbs B, Collin F et al (2003) Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4:249–264. doi:10.1093/biostatistics/4.2.249
Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D (2011) Global cancer statistics. CA Cancer J Clinic 61 (2):69–90. doi:10.3322/caac.20107
Laneve P, Gioia U, Ragno R, Altieri F, Di Franco C, Santini T, Arceci M, Bozzoni I, Caffarelli E (2008) The tumor marker human placental protein 11 is an endoribonuclease. J Biol Chem 283(50):34712–34719. doi:10.1074/jbc.M805759200
Lowy DR, Schiller JT (2006) Prophylactic human papillomavirus vaccines. J Clin Invest 116(5):1167–1173. doi:10.1172/JCI28607
Noordhuis MG, Fehrmann RS, Wisman GB, Nijhuis ER, van Zanden JJ, Moerland PD, Ver Loren van E, Volders HH, Kok M, ten Hoor KA, Hollema H, de Vries EG, de Bock GH, van der Zee AG, Schuuring E (2011) Involvement of the TGF-beta and beta-catenin pathways in pelvic lymph node metastasis in early-stage cervical cancer. Clin Cancer Res 17(6):1317–1330. doi:10.1158/1078-0432.CCR-10-2320
Pyeon D, Ma Newton, Lambert PF, den Boon JA, Sengupta S, Marsit CJ, Woodworth CD, Connor JP, Haugen TH, Smith EM, Kelsey KT, Turek LP, Ahlquist P (2007) Fundamental differences in cell cycle deregulation in human papillomavirus-positive and human papillomavirus-negative head/neck and cervical cancers. Cancer Res 67:4605–4619. doi:10.1158/0008-5472.CAN-06-3619
Rosty C, Sheffer M, Tsafrir D, Stransky N, Tsafrir I, Peter M, de Cremoux P, de La Rochefordiere A, Salmon R, Dorval T, Thiery JP, Couturier J, Radvanyi F, Domany E, Sastre-Garau X (2005) Identification of a proliferation gene cluster associated with HPV E6/E7 expression level and viral DNA load in invasive cervical carcinoma. Oncogene 24(47):7094–7104. doi:10.1038/sj.onc.1208854
Rozanov DV, Savinov AY, Williams R, Liu K, Golubkov VS, Krajewski S, Strongin AY (2008) Molecular signature of MT1-MMP: transactivation of the downstream universal gene network in cancer. Cancer Res 68(11):4086–4096. doi:10.1158/0008-5472.CAN-07-6458
Rudy J, Valafar F (2011) Empirical comparison of cross-platform normalization methods for gene expression data. BMC Bioinform 12(1):467. doi:10.1186/1471-2105-12-467
Scotto L, Narayan G, Nandula SV, Arias-Pulido H, Subramaniyam S, Schneider A, Kaufmann AM, Wright JD, Pothuri B, Mansukhani M, Murty VV (2008) Identification of copy number gain and overexpressed genes on chromosome arm 20q by an integrative genomic approach in cervical cancer: potential role in progression. Genes Chromosom Cancer 47(9):755–765. doi:10.1002/gcc.20577
Slebos RJ, Yi Y, Ely K, Carter J, Evjen A, Zhang X, Shyr Y, Murphy BM, Cmelak AJ, Burkey BB, Netterville JL, Levy S, Yarbrough WG, Chung CH (2006) Gene expression differences associated with human papillomavirus status in head and neck squamous cell carcinoma. Clin Cancer Res 12(3 Pt 1):701–709. doi:10.1158/1078-0432.CCR-05-2017
Subramanian A, Tamayo P, Mootha VK et al (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A 102:15545–15550. doi:10.1073/pnas.0506580102
Ueno M, Itoh M, Kong L, Sugihara K, Asano M, Takakura N (2005) PSF1 is essential for early embryogenesis in mice. Mol Cell Biol 25(23):10528–10532. doi:10.1128/MCB.25.23.10528-10532.2005
Wieland S, Thimme R, Purcell RH, Chisari FV (2004) Genomic analysis of the host response to hepatitis B virus infection. Proc Natl Acad Sci USA 101(17):6669–6674. doi:10.1073/pnas.0401771101
Wilensky RL, Shi Y, Mohler ER 3rd, Hamamdzic D, Burgert ME, Li J, Postle A, Fenning RS, Bollinger JG, Hoffman BE, Pelchovitz DJ, Yang J, Mirabile RC, Webb CL, Zhang L, Zhang P, Gelb MH, Walker MC, Zalewski A, Macphee CH (2008) Inhibition of lipoprotein-associated phospholipase A2 reduces complex coronary atherosclerotic plaque development. Nat Med 14(10):1059–1066. doi:10.1038/nm.1870
Wilson CL, Miller CJ (2005) Simpleaffy: a BioConductor package for Affymetrix Quality Control and data analysis. Bioinformatics 21:3683–3685. doi:10.1093/bioinformatics/bti605
Wolfer A, Ramaswamy S (2011) MYC and metastasis. Cancer Res 71:2034–2037. doi:10.1158/0008-5472.CAN-10-3776
Zhang J, Cao W, Xu Q, Chen WT (2011) The expression of EMP1 is downregulated in oral squamous cell carcinoma and possibly associated with tumour metastasis. J Clin Pathol 64(1):25–29. doi:10.1136/jcp.2010.082404
Acknowledgments
We wish to acknowledge the kind help and insightful correspondence with B. Bachtiary and M. Pintilie, especially M. Pintilie is sincerely thanked for providing the R code of the variance-component analysis. We like to thank Scott Preiss for critical comments on the initial manuscript and for sharing his competence in HPV research.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Statement of translational relevance: At present, there are no generally accepted biomarkers for cervical cancer available. Here, we utilize an already approved approach for biomarker discovery on a large-scale pooled microarray dataset. Eleven highly reliable probes resulting from this analysis feature cervical cancer biomarker properties. Moreover, overlap between our gene signatures and those from studies available in the literature affirm the utility of this approach. However, the most striking findings are that the gene expression signatures were associated with signatures observed for patients treated using angiocidin and darapladib. Previous studies have shown that angiocidin induces differentiation of monocytes to macrophages and induce anti-tumoral action, whereas darapladib exerts anti-inflammatory action. Therefore, we suggest these two agents as potential candidates for novel treatment options in cervical cancer patients.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Koch, M., Wiese, M. Gene expression signatures of angiocidin and darapladib treatment connect to therapy options in cervical cancer. J Cancer Res Clin Oncol 139, 259–267 (2013). https://doi.org/10.1007/s00432-012-1317-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-012-1317-9