Abstract
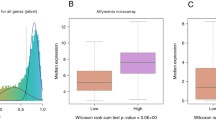
Glioblastoma is the most common and deadly type of brain cancer. Over the past decade, several divergent genetic pathways have been implicated in the initiation, progression and clinical outcome of this disease. As our understanding of GBM expands and identifies actionable targets specific to individual tumors, there will be a pressing need for the development of new tools that will maximize the use of limited clinical samples to enable the employment of personalized care paradigms. We used PrimePCR validated assays to generate a custom real-time PCR screening panel, containing 74 previously published mRNA targets showing gene expression changes in glioblastoma, and five house-keeping genes. A cohort of 19 frozen brain specimens were analyzed, including WHO Grade II oligodendroglioma (n = 3), WHO Grade II astrocytoma (n = 2), WHO Grade III astrocytoma (n = 1), and glioblastoma (n = 13). Four normal brain samples were also analyzed. We performed RNA extraction, followed by cDNA synthesis, multiplexed pre-amplification and SYBR-based qPCR, to generate expression profiles on all samples. We demonstrated that the workflow shows high tolerance to variation in RNA quality (RIN 8.5-4) and high sensitivity in detection. cDNA input that is equivalent to 3 ng of starting RNA was sufficient to conduct accurate semiquantitative analysis of the panel of 79 assays. Using principal component analysis, we were able to accurately separate glioblastoma from low-grade glioma. The two WHO Grade III tumors analyzed clustered with glioblastoma, but showed more similarity to Grade II gliomas. In this study, we have shown the feasibility of consolidating high-throughput data into a single functional panel capable of accurately classifying glioma specimens based solely on semiquantitative gene expression profiling.



Similar content being viewed by others
References
CBTRUS (2005) Statistical report: primary brain tumours in the United States, 1998–2002. Hinsdale: Central Brain Tumour Registry of the United States
Zhang X, Zhang W, Cao WD, Cheng G, Zhang YQ (2012) Glioblastoma multiforme: molecular characterization and current treatment strategy. Exp Ther Med 3(1):9–14
Kleihues P, Cavenee WK (eds) (2000) Pathology and genetics of tumours of the nervous system. IARC Press, Lyon
Giannini C, Scheithauer BW, Weaver AL et al (2001) Oligodendrogliomas: reproducibility and prognostic value of histologic diagnosis and grading. J Neuropathol Exp Neurol 60(3):248–262
Rickman DS, Bobek MP, Misek DE et al (2001) Distinctive molecular profiles of high-grade and low-grade gliomas based on oligonucleotide microarray analysis. Cancer Res 61(18):6885–6891
McLendon R, Friedman A, Bigner D et al (2008) Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 455(7216):1061–1068
Frattini V, Trifonov V, Chan JM et al (2013) The integrated landscape of driver genomic alterations in glioblastoma. Nat Genet 45(10):1141–1149
Parsons DW, Jones S, Zhang X et al (2008) An integrated genomic analysis of human glioblastoma multiforme. Science 321(5897):1807–1812
Verhaak RG, Hoadley KA, Purdom E et al (2010) Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 17(1):98–110
Kibschull M, Lye SJ, Okino ST, Sarras H (2015) Quantitative large scale gene expression profiling from human stem cell culture micro samples using multiplex pre-amplification. Syst Biol Reprod Med 3:1–8
Acknowledgments
We thank Dr. Yan Wang and Dr. Steven Okino for critically reading the manuscript. We wish to acknowledge the Labatt Brain Tumour Research Centre Tumour and Tissue Repository which is supported by b.r.a.i.nchild and Meagan’s Walk. This work was supported by grants from b.r.a.i.n.child and the American College of Surgeons Martin Research Fellowship.
Author information
Authors and Affiliations
Corresponding author
Additional information
Haya Sarras and Megan Wu have contributed equally to the manuscript.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Sarras, H., Wu, M., Celebre, A. et al. A novel amplification-based approach to enable gene expression profiling from small clinical tumor specimens. J Neurooncol 126, 69–75 (2016). https://doi.org/10.1007/s11060-015-1953-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11060-015-1953-4