Abstract
It is generally believed that cerebellar granule neurons originate exclusively from granule neuron precursors (GNPs) in the external germinal layer (EGL). Here we identified a rare population of neuronal progenitors in mouse developing cerebellum that expresses Nestin. Although Nestin is widely considered a marker for multipotent stem cells, these Nestin-expressing progenitors (NEPs) are committed to the granule neuron lineage. Unlike conventional GNPs, which reside in the outer EGL and proliferate extensively, NEPs reside in the deep part of the EGL and are quiescent. Expression profiling revealed that NEPs are distinct from GNPs and, in particular, express markedly reduced levels of genes associated with DNA repair. Consistent with this, upon aberrant activation of Sonic hedgehog (Shh) signaling, NEPs exhibited more severe genomic instability and gave rise to tumors more efficiently than GNPs. These studies revealed a previously unidentified progenitor for cerebellar granule neurons and a cell of origin for medulloblastoma.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Lendahl, U., Zimmerman, L.B. & McKay, R.D. CNS stem cells express a new class of intermediate filament protein. Cell 60, 585–595 (1990).
Morshead, C.M. et al. Neural stem cells in the adult mammalian forebrain: a relatively quiescent subpopulation of subependymal cells. Neuron 13, 1071–1082 (1994).
Shi, H. et al. Nestin expression defines both glial and neuronal progenitors in postnatal sympathetic ganglia. J. Comp. Neurol. 508, 867–878 (2008).
Vukojevic, K., Petrovic, D. & Saraga-Babic, M. Nestin expression in glial and neuronal progenitors of the developing human spinal ganglia. Gene Expr. Patterns 10, 144–151 (2010).
Hyder, C.L., Isoniemi, K.O., Torvaldson, E.S. & Eriksson, J.E. Insights into intermediate filament regulation from development to ageing. J. Cell Sci. 124, 1363–1372 (2011).
Lee, A. et al. Isolation of neural stem cells from the postnatal cerebellum. Nat. Neurosci. 8, 723–729 (2005).
Yang, Z.J. et al. Medulloblastoma can be initiated by deletion of Patched in lineage-restricted progenitors or stem cells. Cancer Cell 14, 135–145 (2008).
Sutter, R. et al. Cerebellar stem cells act as medulloblastoma-initiating cells in a mouse model and a neural stem cell signature characterizes a subset of human medulloblastomas. Oncogene 29, 1845–1856 (2010).
Sotelo, C., Alvarado-Mallart, R.M., Frain, M. & Vernet, M. Molecular plasticity of adult Bergmann fibers is associated with radial migration of grafted Purkinje cells. J. Neurosci. 14, 124–133 (1994).
Alder, J., Cho, N.K. & Hatten, M.E. Embryonic precursor cells from the rhombic lip are specified to a cerebellar granule neuron identity. Neuron 17, 389–399 (1996).
Rao, G., Pedone, C.A., Coffin, C.M., Holland, E.C. & Fults, D.W. c-Myc enhances sonic hedgehog-induced medulloblastoma formation from nestin-expressing neural progenitors in mice. Neoplasia 5, 198–204 (2003).
Rao, G. et al. Sonic hedgehog and insulin-like growth factor signaling synergize to induce medulloblastoma formation from nestin-expressing neural progenitors in mice. Oncogene 23, 6156–6162 (2004).
Tanori, M. et al. Developmental and oncogenic effects of insulin-like growth factor-I in Ptc1+/− mouse cerebellum. Mol. Cancer 9, 53 (2010).
Frappart, P.O., Lee, Y., Lamont, J. & McKinnon, P.J. BRCA2 is required for neurogenesis and suppression of medulloblastoma. EMBO J. 26, 2732–2742 (2007).
Mills, J. et al. Critical role of integrin-linked kinase in granule cell precursor proliferation and cerebellar development. J. Neurosci. 26, 830–840 (2006).
Encinas, J.M., Vaahtokari, A. & Enikolopov, G. Fluoxetine targets early progenitor cells in the adult brain. Proc. Natl. Acad. Sci. USA 103, 8233–8238 (2006).
Ben-Arie, N. et al. Math1 is essential for genesis of cerebellar granule neurons. Nature 390, 169–172 (1997).
Lumpkin, E.A. et al. Math1-driven GFP expression in the developing nervous system of transgenic mice. Gene Expr. Patterns 3, 389–395 (2003).
Uchida, N. et al. Direct isolation of human central nervous system stem cells. Proc. Natl. Acad. Sci. USA 97, 14720–14725 (2000).
Reynolds, B.A. & Weiss, S. Generation of neurons and astrocytes from isolated cells of the adult mammalian central nervous system. Science 255, 1707–1710 (1992).
Vintersten, K. et al. Mouse in red: red fluorescent protein expression in mouse ES cells, embryos, and adult animals. Genesis 40, 241–246 (2004).
Aruga, J. et al. A novel zinc finger protein, zic, is involved in neurogenesis, especially in the cell lineage of cerebellar granule cells. J. Neurochem. 63, 1880–1890 (1994).
Balordi, F. & Fishell, G. Mosaic removal of hedgehog signaling in the adult SVZ reveals that the residual wild-type stem cells have a limited capacity for self-renewal. J. Neurosci. 27, 14248–14259 (2007).
Mao, X., Fujiwara, Y., Chapdelaine, A., Yang, H. & Orkin, S.H. Activation of EGFP expression by Cre-mediated excision in a new ROSA26 reporter mouse strain. Blood 97, 324–326 (2001).
Machold, R. & Fishell, G. Math1 is expressed in temporally discrete pools of cerebellar rhombic-lip neural progenitors. Neuron 48, 17–24 (2005).
Wechsler-Reya, R.J. & Scott, M.P. Control of neuronal precursor proliferation in the cerebellum by Sonic Hedgehog. Neuron 22, 103–114 (1999).
Kenney, A.M., Cole, M.D. & Rowitch, D.H. Nmyc upregulation by sonic hedgehog signaling promotes proliferation in developing cerebellar granule neuron precursors. Development 130, 15–28 (2003).
Orford, K.W. & Scadden, D.T. Deconstructing stem cell self-renewal: genetic insights into cell-cycle regulation. Nat. Rev. Genet. 9, 115–128 (2008).
Lombard, D.B. et al. DNA repair, genome stability, and aging. Cell 120, 497–512 (2005).
Burhans, W.C. & Weinberger, M. DNA replication stress, genome instability and aging. Nucleic Acids Res. 35, 7545–7556 (2007).
Ellis, T. et al. Patched 1 conditional null allele in mice. Genesis 36, 158–161 (2003).
Ahn, S. & Joyner, A.L. In vivo analysis of quiescent adult neural stem cells responding to Sonic hedgehog. Nature 437, 894–897 (2005).
Balordi, F. & Fishell, G. Hedgehog signaling in the subventricular zone is required for both the maintenance of stem cells and the migration of newborn neurons. J. Neurosci. 27, 5936–5947 (2007).
Aguilera, A. & Gomez-Gonzalez, B. Genome instability: a mechanistic view of its causes and consequences. Nat. Rev. Genet. 9, 204–217 (2008).
Frappart, P.O. et al. Recurrent genomic alterations characterize medulloblastoma arising from DNA double-strand break repair deficiency. Proc. Natl. Acad. Sci. USA 106, 1880–1885 (2009).
Silbereis, J. et al. Astroglial cells in the external granular layer are precursors of cerebellar granule neurons in neonates. Mol. Cell. Neurosci. 44, 362–373 (2010).
Nicot, A., Lelievre, V., Tam, J., Waschek, J.A. & DiCicco-Bloom, E. Pituitary adenylate cyclase-activating polypeptide and sonic hedgehog interact to control cerebellar granule precursor cell proliferation. J. Neurosci. 22, 9244–9254 (2002).
Rios, I., Alvarez-Rodriguez, R., Marti, E. & Pons, S. Bmp2 antagonizes sonic hedgehog-mediated proliferation of cerebellar granule neurones through Smad5 signalling. Development 131, 3159–3168 (2004).
Fogarty, M.P., Kessler, J.D. & Wechsler-Reya, R.J. Morphing into cancer: the role of developmental signaling pathways in brain tumor formation. J. Neurobiol. 64, 458–475 (2005).
Pons, S., Trejo, J.L., Martinez-Morales, J.R. & Marti, E. Vitronectin regulates Sonic hedgehog activity during cerebellum development through CREB phosphorylation. Development 128, 1481–1492 (2001).
Schuller, U. et al. Acquisition of granule neuron precursor identity is a critical determinant of progenitor cell competence to form Shh-induced medulloblastoma. Cancer Cell 14, 123–134 (2008).
Kaye, J.A. et al. DNA breaks promote genomic instability by impeding proper chromosome segregation. Curr. Biol. 14, 2096–2106 (2004).
Jasin, M. Chromosome breaks and genomic instability. Cancer Invest. 18, 78–86 (2000).
Oliver, T.G. et al. Loss of patched and disruption of granule cell development in a pre-neoplastic stage of medulloblastoma. Development 132, 2425–2439 (2005).
Pazzaglia, S. et al. High incidence of medulloblastoma following X-ray-irradiation of newborn Ptc1 heterozygous mice. Oncogene 21, 7580–7584 (2002).
Pazzaglia, S. et al. The genetic control of chemically and radiation-induced skin tumorigenesis: a study with carcinogenesis-susceptible and carcinogenesis-resistant mice. Radiat. Res. 158, 78–83 (2002).
Lee, Y. & McKinnon, P.J. DNA ligase IV suppresses medulloblastoma formation. Cancer Res. 62, 6395–6399 (2002).
Fernandez, L.A. et al. Oncogenic YAP promotes radioresistance and genomic instability in medulloblastoma through IGF2-mediated Akt activation. Oncogene 31, 1923–1937 (2012).
Beer, S. et al. Developmental context determines latency of MYC-induced tumorigenesis. PLoS Biol. 2, e332 (2004).
Acknowledgements
We thank J. Oesterling for flow cytometric analysis; Z. Liu, J. Pei and J. Testa for cytogenetic analysis; A. Efimov for microscopy analysis; Q. Cai for histological analysis; R. Segal (Dana Farber Cancer Institute) for Zic1 antibody; and D. Wiest, F. Roegiers and T. Yen for helpful discussions. This research was supported by the W.W. Smith Charitable Trust (Z.Y.), a generous gift from Cathie and Pete Getchell (Z.Y.), a US National Institutes of Health Postdoctoral training grant (5T32CA009035-37, L.W.Y.), grants from the US National Cancer institute (R01-CA178380, Z.Y.; R01-CA122759, R.J.W.-R.), pilot funding from the US National Institutes of Health (U19-AI067798, R.J.W.-R.) and the California Institute for Regenerative Medicine (LA1-01747, R.J.W.-R.).
Author information
Authors and Affiliations
Contributions
Z.Y. and R.J.W.-R. conceived the project. P.L., F.D., L.W.Y., T.L., R.E.M. and R.T. performed the experiments. Z.Y., P.L., J.W., A.B. and R.J.W.-R. analyzed the data. G.E. provided reagents. Z.Y. prepared the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Nestin expression in the developing cerebellum.
Cerebellar sections from Nestin-CFP animals at various stages were immunostained with indicated antibodies. (a) At E14.5, no CFP+ cells were present in the EGL (arrows) and most CFP+ cells were localized in the ventricular zone (arrowheads). (b) Higher magnification of boxed region in figure a reveals the absence of CFP+ cells in the rhombic lip (arrows). (c) At E16.5, NEPs (CFP+, arrows) were found in the deep part of the EGL and NSCs (Sox2+, arrow heads) in the ventricular zone. (d) Higher magnification of boxed region in figure b shows the localization of NEPs in the EGL (arrows). (e) At P21, only astroglial cells (S100+, arrows) still express CFP. Scale bar: a, c and e (200 μm); b and d (80 μm).
Supplementary Figure 2 Nestin expressing cells in the cerebellar white matter.
(a) Cerebellar slices were prepared from Math1-GFP/Nestin-CFP animals at P4. The white matter was dissected under a fluorescent microscope (as shown by the dotted yellow line). (b) The dissected white matter was collected for cell dissociation. (c) Cells isolated from the white matter of P4 Math1-GFP/Nestin-CFP animals, were analyzed by flow cytometry for expression of GFP and CFP. Approximately 18% of cells in the white matter were positive for Nestin-CFP. (d and e) Nestin-CFP positive cells in the white matter were stained with isotype control (mouse IgG) or anti-Prominin1 prior to FACs analysis. ∼30% of CFP positive cells were Prominin1+, suggesting that Nestin-expressing cells in the cerebellar white matter at P4 include NSCs. Scale bar: a (400μm); b (2mm).
Supplementary Figure 3 Expression of Nestin and Math1 in purified CFP+ and GFP+ cells.
(a-d) Immediately after being isolated from the EGL of Math1-GFP/Nestin-CFP animals at P4, GFP+ cells (a) were stained for Math1 (red, b). Merged image of a and b indicates that all GFP+ cells express Math1 (c). GFP+ cells were also immunostained for Nestin (red, d). No Nestin expression was found among GFP+ cells. (e-g) CFP+ cells (e) from the EGL of Math1-GFP/Nestin-CFP cerebellum at P4 were immunostained for Nestin (red, f). The merged image of e and f shows Nestin expression in all purified CFP+ cells (g). CFP+ cells were immunostained for Math1 (red, h). No Math1+ cells were detected among CFP+ cell population. Scale bar: 200μm.
Supplementary Figure 4 Differentiation of purified NSCs.
NSCs (Prominin1+, Lin- cells) isolated from P4 Nestin-CFP/Math1-GFP cerebellum, were cultured in vitro for 3 days and immunostained for neurons (ß-tubulin+, a), Bergmann glia (S100ß+, b) and oligodendrocytes (O4+, c). Scale bar: 60μm.
Supplementary Figure 5 Fibers of Bergmann glia remaining on cerebellar surface of Nestin-CreERT2–R26R-GFP cerebellum.
Cerebellar sections prepared from Nestin-CreERT2/R26R-GFP mouse at P21 after tamoxifen treatment at P4 were stained for GFP (green, a and b), S100ß (red, a), and counterstained with DAPI (blue, b). GFP+ fibers on the cerebellar surface were positive for S100ß, and negative for DAPI (arrows). Scale bar: 67 μm.
Supplementary Figure 6 DNA instability in proliferating NEPs.
NEPs and GNPs purified from Math1-GFP/Nestin-CFP cerebella at P4, were treated with recombinant Shh in vitro for 48hrs. (a) Proliferating NEPs and GNPs were then harvested to examine the expression of Chek1, Lig3 and Parp1 by quantitative PCR. Expression of all genes in NEPs is normalized to their relative expression in GNPs. (b) PCR products were examined by gel electrophoresis. (c) Percentage of BrdU incorporated GNPs and NEPs among Cre-infected cells. (d) The table summarizes the number of metaphases with chromosomal alterastions in metaphase spread from Ptch1 deficient GNPs and NEPs. More chromosomal abnormalities were detected among NEPs compared with GNPs (Chi-square test, χ2=7.05, P=0.00793, n=50 for GNPs and n=54 for NEPs.) Data in a and c represent means of triplicate experiments ±SEM and significance determined with two-tailed Student's t test <0.001, P<0.01. (a) Chek1 of NEPs vs GNPs, P=0.00085; Lig3 of NEPs vs GNPs, P=0.0171; Parp1 of NEPs vs GNPs, P=0.0106; (b) P=0.837
Supplementary Figure 7 The similar genetic profile of NEP- and GNP-derived tumor cells.
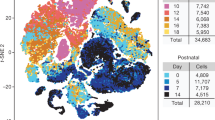
MB cells were isolated from Math1-CreERT2/Ptch1C/C mice and Nestin-CreERT2/Ptch1C/C mice, and total RNAs were extracted from tumor cells for microarray analysis. Genetic profiles of tumor cells were compared with the normal cerebella (downloaded from the NCBI Gene Expression Omnibus (www.ncbi.nlm.nih.gov/geo) with the accession number GSE11859) by PCA.
Supplementary Figure 8 The tumorigenicity of transplanted NEPs and GNPs.
(a) Two different primer sets were designed to analyze Ptch1 genomic DNA. Primer set “a” amplifies part of exon 3, which is deleted in the mutant allele; primer set “b” is specific for exon 14, which is present in both wild type and mutant alleles. (b) After tamoxifen treatment at P4, GNPs and NEPs were purified from the EGL of Math1-GFP/Math1-CreERT2/Ptch1C/C cerebellum and Nestin-CFP/Nestin-CreERT2/Ptch1C/C cerebellum at P8, respectively. GNPs from wild type cerebellum at P8 and tumor cells from Math1-Cre/Ptch1C/C at 8 weeks of age were isolated as controls. Genomic DNA extracted from those cells was used for quantitative PCR using primer sets a and b. (c) The amount of undeleted Ptch1 in each sample was calculated by dividing the level of exon 3 product by the total amount of Ptch1 DNA (represented by exon 14 product). (d) The table shows the number of animals with tumors from the different amount of cells following the transplantation. Data in c represent means of triplicate experiments ± SEM. and significance determined with two-tailed Student's t test. (Ptch1 deletion among NEPs vs GNPs, P=0.486).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Table 1 (PDF 5083 kb)
Rights and permissions
About this article
Cite this article
Li, P., Du, F., Yuelling, L. et al. A population of Nestin-expressing progenitors in the cerebellum exhibits increased tumorigenicity. Nat Neurosci 16, 1737–1744 (2013). https://doi.org/10.1038/nn.3553
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.3553
This article is cited by
-
Nestin is essential for cellular redox homeostasis and gastric cancer metastasis through the mediation of the Keap1–Nrf2 axis
Cancer Cell International (2021)
-
Depletion of kinesin motor KIF20A to target cell fate control suppresses medulloblastoma tumour growth
Communications Biology (2021)
-
Nestin regulates cellular redox homeostasis in lung cancer through the Keap1–Nrf2 feedback loop
Nature Communications (2019)
-
Impaired Cerebellar Development in Mice Overexpressing VGF
Neurochemical Research (2019)
-
Childhood cerebellar tumours mirror conserved fetal transcriptional programs
Nature (2019)